Peptide Sequencing Services
Peptides are composed of two or more amino acids linked together by peptide bonds formed through dehydration condensation. Peptides containing fewer than 10 amino acids are referred to as oligopeptides (small peptides), those with more than 10 are classified as polypeptides, and those exceeding 50 amino acids are considered proteins. Peptides play a wide range of physiological roles in organisms, participating in cellular signal transduction, regulating physiological processes, and acting as hormones.
The amino acid composition and sequence of peptides are crucial in peptide identification, providing valuable reference information for determining peptide types. Peptide sequencing services can analyze the amino acid composition and sequence of peptides. The most commonly used methods for peptide sequencing are Edman degradation and mass spectrometry, which are based on distinct chemical and physical principles. For example, Edman degradation identifies the entire sequence by sequentially removing and identifying the N-terminal amino acids of a polypeptide. On the other hand, mass spectrometry determines the molecular weight and amino acid composition of peptides by analyzing their ions' motion in electric and magnetic fields based on their mass-to-charge ratio (m/z).
Services at MtoZ Biolabs
Leveraging advanced mass spectrometry platforms, fully automated protein/peptide sequencers, and high-performance liquid chromatography systems, MtoZ Biolabs offers professional and accurate peptide sequencing services for both individual and mixed peptides.
1. Mass Spectrometry
Mass spectrometry is one of the most widely used methods for peptide sequencing, particularly for mixed peptide sequencing. It includes de novo peptide sequencing and database-based peptide sequencing.The standard workflow of mass spectrometry-based peptide sequencing services involve liquid chromatography separation of peptide samples followed by ionization and introduction into a mass analyzer. The ions are separated and detected based on their mass-to-charge ratios, generating mass spectrometry data. This data is then processed using leading de novo peptide analysis software such as PEAKS Studio to decode the complete amino acid composition of the peptide sequences. Alternatively, database search algorithms can be used to compare the obtained peptide data with known sequences in databases, identifying matching peptide sequences.
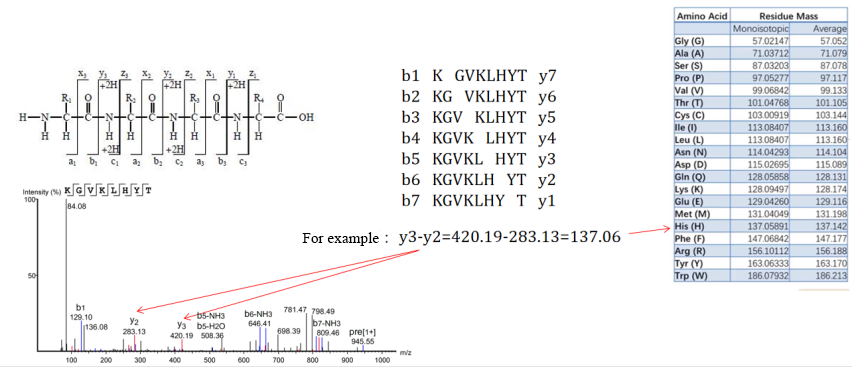
Figure 1. Schematic Diagram of De Novo Peptide Sequencing.
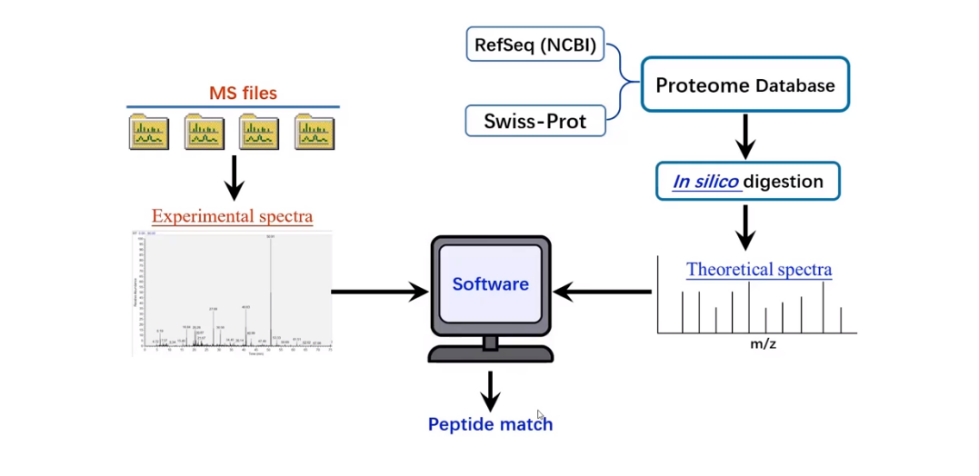
Figure 2. Schematic Diagram of Peptide Sequencing Based on Database Matching.
2. Edman Degradation Method
The Edman degradation method is a classical technique for peptide sequencing, primarily suitable for sequencing single peptides. This method requires high sample purity and is not applicable for peptides with an N-terminal blockage or modifications in the sequence. It utilizes phenylisothiocyanate (PITC) to react with the N-terminal amino acid of the peptide, forming a phenylthiocarbamoyl (PTC) derivative. Under acidic conditions, the PTC derivative undergoes cyclization, releasing the N-terminal amino acid as a thiazolinone derivative. This derivative is then identified using chromatography or mass spectrometry. By repeating this process, the N-terminal amino acids of the peptide are sequentially removed and identified, ultimately determining the complete peptide sequence.
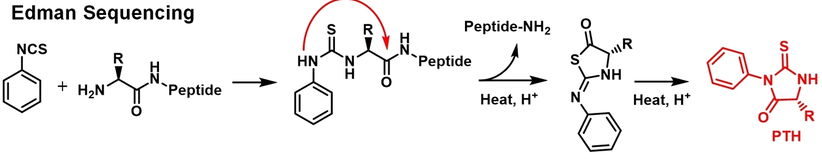
Shi, Q. et al. Angew Chem Int Ed Engl. 2024.
Figure 3. Schematic Diagram of The Edman Degradation Method.
Service Advantages
1.Advanced Analytical Platforms
MtoZ Biolabs is equipped with multiple state-of-the-art mass spectrometers, including Orbitrap Astral, Orbitrap Exploris 480, Orbitrap Fusion™ Lumos™ Tribrid™, and Q Exactive, as well as advanced fully automated protein/peptide sequencers(PPSQ33A).
2.Expert Team
Our experienced technical team provides one-on-one support to address all technical inquiries.
3.High-Quality Data
Deep data coverage and strict data quality control. We have a standardized laboratory management system to ensure the accuracy and stability of the experimental process.
Sample Submission Suggestions
1. Sample Types
(1) Single or mixed peptide samples are acceptable.
(2) Liquid/solid samples, cells, bacteria, tissues, blood, urine, paraffin-embedded tissue sections, immune peptides, etc.
Note: For special requirements or sample preparation assistance, please contact us.
Applications
1. Drug Design and Optimization
Peptide sequencing is crucial for understanding drug mechanisms. By analyzing the peptide sequence of drug targets, researchers can design and optimize drug molecules to enhance efficacy and minimize side effects.
2. Biological Product Quality Control
During the production of biopharmaceuticals, peptide sequencing services can monitor product quality and ensure consistency in amino acid sequences, maintaining drug stability and potency.
3. Development of Novel Bioproducts
Peptide sequencing technology has been applied to the development of innovative bioproducts such as antibody drugs, cell therapies, and gene therapies.
4. Tumor Neoantigen Identification
In the process of discovering tumor neoantigens, peptide sequencing services can help determine the tumor neoantigens produced by patient tumor cells and their corresponding antigen peptides, which is of great significance for the development of tumor vaccines.
5. Drug Safety Evaluation
De novo peptide sequencing can help assess drug safety. By detecting variations or mutations in peptide sequences, researchers can identify potential side effects and drug interactions, facilitating safety evaluations.
Case Study
1. Identification and Stability Evaluation of a Novel ACE-Inhibitory Hexapeptide from Camellia Seed Cake
In studies on ACE inhibitory activity under neutral protease, alkaline protease, papain, and trypsin hydrolysis, the Val-Val-Val-Pro-Gln-Asn(VVVPQN) can be obtained by peptide sequencing services based on mass spectrometry. VVVPQN exhibited ACE inhibitory activity with an IC50 value of 0.13 mg/mL (198.66 µmol/L) and inhibited ACE in a non-competitive manner. Molecular docking revealed that VVVPQN formed eight hydrogen bonds with ACE. Stability study results show that VVVPQN maintains high ACE inhibitory activity in weakly acidic and neutral environments. Heat treatment (20-80°C) and Na+, Mg2+ and Fe3+ metal ions have little effect on the activity of VVVPQN.Furthermore, VVVPQN remained relatively stable after simulated gastrointestinal digestion. These findings suggest that VVVPQN, identified in Camellia seed cake, has potential applications in functional foods or antihypertensive drugs.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment (project-dependent)
4.Data Analysis, Peptide Sequence List and Project Report
5. Raw Data Files
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details!
How to order?







