Post-translational Modifications (PTMs) Service
- Oxidation (e.g., methionine sulfoxidation)
- Nitration (e.g., tyrosine nitration)
- Deamidation
- Lipidation
Post-translational modifications play a pivotal role in modulating protein function, localization, stability, and interactions, significantly influencing cellular processes such as signal transduction, gene expression regulation, and disease progression. PTMs encompass a wide variety of chemical modifications, including phosphorylation, acetylation, ubiquitination, methylation, glycosylation, and SUMOylation, each with distinct biological implications.
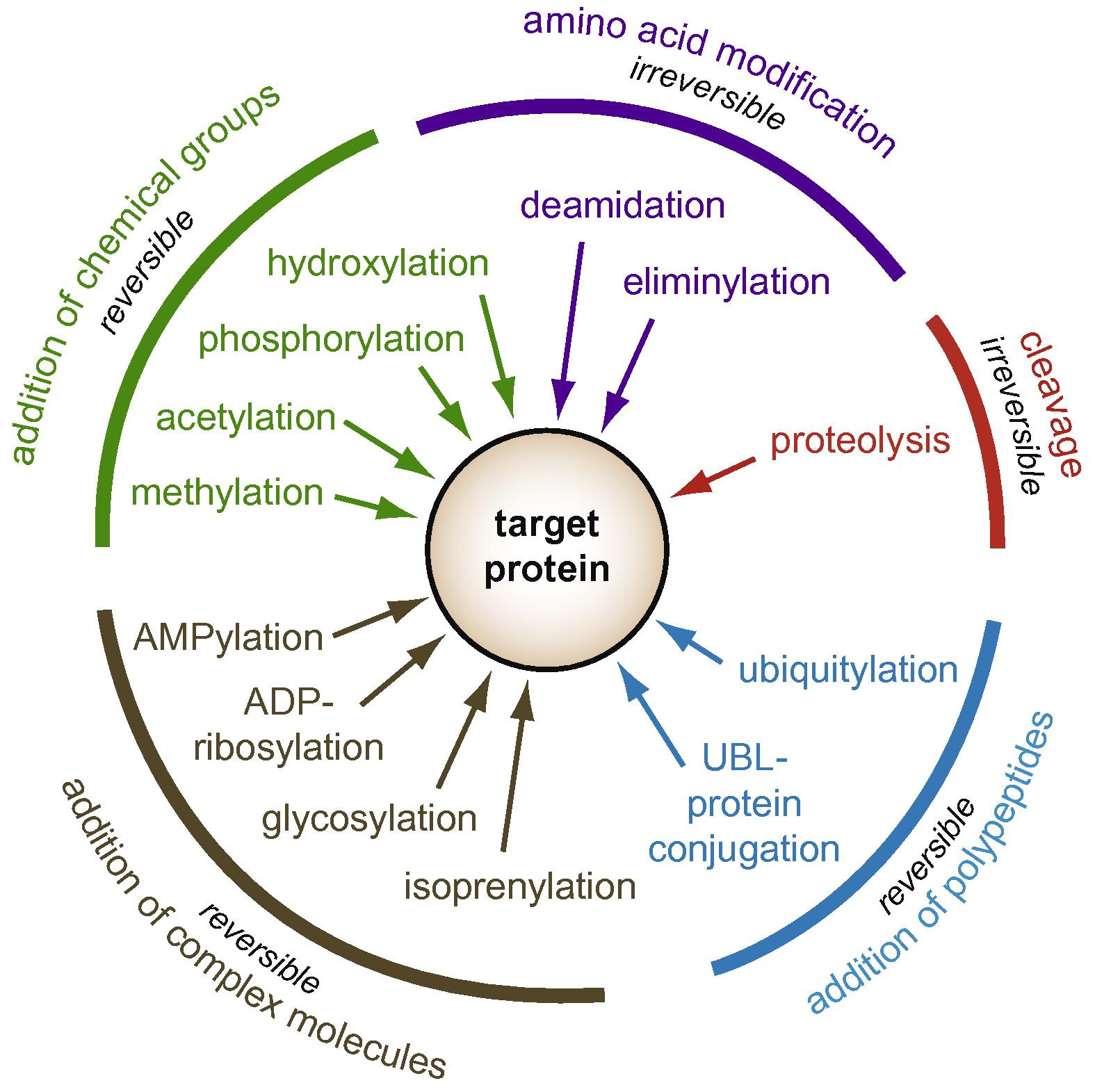
Ribet, D. et al. FEBS Lett. 2010.
Figure1. Diversity of Post-Translational Modifications
Why Analyze PTMs?
Studying PTMs is essential for understanding protein function and regulation. The addition or removal of functional groups or structural modifications can:
Regulate enzyme activity and protein-protein interactions
Mediate cellular signaling pathways
Modulate transcriptional and epigenetic mechanisms
Influence protein stability and degradation (e.g., ubiquitination, SUMOylation)
Determine subcellular localization and trafficking
By leveraging MtoZ Biolabs’ advanced post-translational modifications (PTMs) service, researchers can uncover novel molecular mechanisms, identify disease biomarkers, and accelerate drug discovery efforts.
Service at MtoZ Biolabs
At MtoZ Biolabs, we provide comprehensive post-translational modifications (PTMs) service using cutting-edge mass spectrometry (MS) platforms, including Thermo Fisher Orbitrap Fusion Lumos, Q Exactive HF, and FTICR MS, combined with advanced liquid chromatography (LC) and enrichment techniques. Our high-resolution, high-sensitivity LC-MS/MS workflows ensure precise identification, site localization, and relative quantification of PTMs, empowering researchers to elucidate protein modifications with exceptional accuracy.
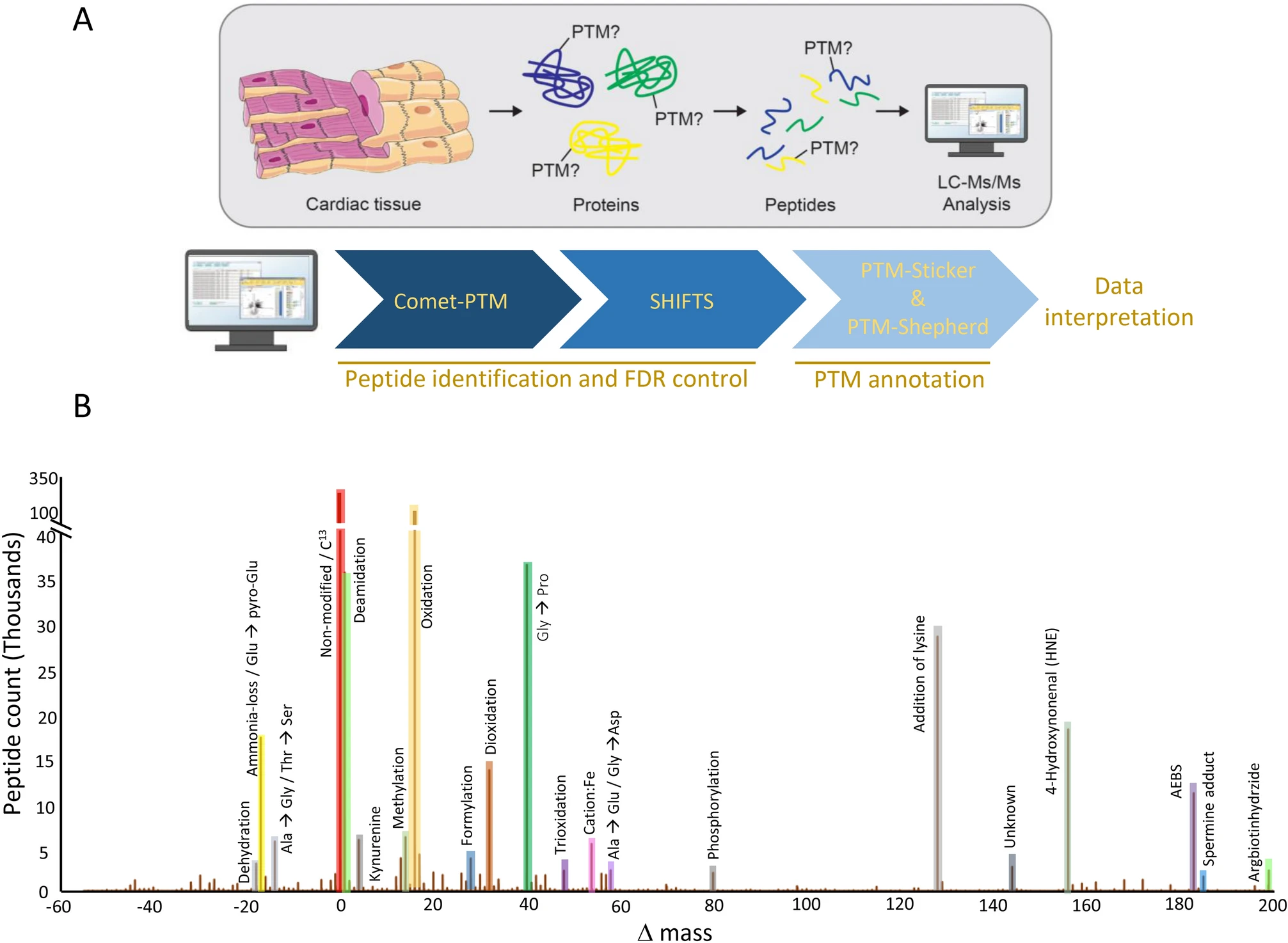
Bagwan, N. et al. Sci Rep. 2021.
Figure2. Shotgun Proteomics Workflow and Analysis Pipeline used for PTM Identifications
We offer a broad spectrum of post-translational modifications (PTMs) service covering both common and rare modifications. Our service workflow ensures accurate and reproducible identification, mapping, and quantification of PTMs with minimal sample requirements.
1. PTM Identification and Site Mapping
Using high-resolution LC-MS/MS, we precisely map modification sites on target proteins. Our PTM site localization algorithms leverage database-dependent and de novo sequencing approaches, ensuring high-confidence PTM assignments.
2. Quantitative PTM Analysis
For relative and absolute quantification of PTMs, we utilize:
√ Label-Free Quantification (LFQ) – Suitable for large-scale PTM profiling
√ Stable Isotope Labeling (SILAC, TMT, iTRAQ) – Enables accurate comparative PTM analysis across different conditions
√ Parallel Reaction Monitoring (PRM) and Selected Reaction Monitoring (SRM) – Highly targeted quantification for specific PTMs
3. Enrichment Strategies for Low-Abundance PTMs
Many PTMs occur at low stoichiometry, making enrichment a crucial step before LC-MS/MS analysis. We offer highly specific enrichment strategies, including:
√ Phosphopeptide Enrichment (TiO₂, Fe-NTA, IMAC, MOAC)
√ Ubiquitinated Peptide Enrichment (Tandem Ubiquitin Binding Entities, TUBEs)
√ Glycopeptide Enrichment (Lectin-based and HILIC methods)
√ Acetylation, Methylation, and SUMOylation Affinity-Based Enrichment
4. PTM-Specific Antibody-Based Methods
For targeted PTM validation and screening, we employ Western blot, ELISA, and IP-MS approaches. These complementary methods enhance the reliability of our LC-MS/MS findings.
PTM Types We Analyze
MtoZ Biolabs offers comprehensive PTM characterization for a wide range of modifications:
1. Phosphorylation Analysis
Phosphorylation is one of the most well-studied PTMs, regulating numerous cell signaling pathways, enzyme activity, and protein function. Our phosphoproteomics service utilizes IMAC, TiO₂, and Fe-NTA enrichment combined with high-resolution MS to pinpoint phosphorylation sites with high specificity and sensitivity.
2. Acetylation and Methylation Analysis
Histone and non-histone protein modifications via acetylation/methylation impact chromatin remodeling, gene expression, and protein-protein interactions. We provide site-specific mapping and quantification of these epigenetic modifications.
3. Ubiquitination and SUMOylation Analysis
Ubiquitination and SUMOylation play key roles in protein degradation, DNA repair, and cell cycle regulation. Due to their transient nature, our advanced affinity-based enrichment strategies coupled with LC-MS/MS analysis ensure reliable identification.
4. Glycosylation Analysis
Glycosylation affects protein folding, stability, and receptor-ligand interactions. We offer N-glycan and O-glycan profiling using enzymatic digestion, HILIC-based enrichment, and MS/MS fragmentation techniques to characterize glycosylation patterns.
5. Oxidation, Nitration, and Other Rare PTMs
We also analyze less common modifications such as:
Why Choose MtoZ Biolabs?
✅ State-of-the-Art Mass Spectrometry Platform – High-resolution MS ensures superior PTM characterization.
✅Customized Experimental Design – Tailored workflows to match research objectives.
✅Advanced Bioinformatics Analysis – Industry-leading PTM identification and quantification pipelines.
✅Comprehensive Service Portfolio – PTM analysis integrated with protein interaction studies, structural proteomics, and metabolomics.
✅Expert Consultation – PTM specialists provide in-depth project guidance.
Sample Submission Suggestions
To ensure high-quality results, we recommend submitting samples under the following guidelines:
✔ Protein Samples: ≥100 µg total protein per sample (cell lysates, tissue extracts, purified proteins)
✔ Peptide Samples: ≥10 µg digested peptides (trypsin/LysC)
✔ Biological Replicates: Minimum 3 replicates per condition for quantitative studies
✔ Enrichment Kits: If available, specify PTM-specific antibodies or affinity-based reagents used for pre-enrichment
MtoZ Biolabs provides detailed sample submission guidelines and pre-experiment consultation to optimize study design.
Case Study
The heart is a high-energy-consuming organ involved in various physiological functions, and the features of PTMs in its proteome, as well as changes in these modifications during diseases, have not been fully studied. This study employed large-scale proteomics analysis techniques, utilizing mass spectrometry (LC-MS/MS) to analyze human heart samples. Through various separation techniques and data analysis methods, combined with strategies like affinity enrichment and chemical modification labeling, the study comprehensively identified a variety of PTMs (such as phosphorylation, acetylation, ubiquitination, etc.) in the heart. Over 100,000 PTM sites were identified in the human heart, covering a wide range of modification types, including phosphorylation, acetylation, and methylation.
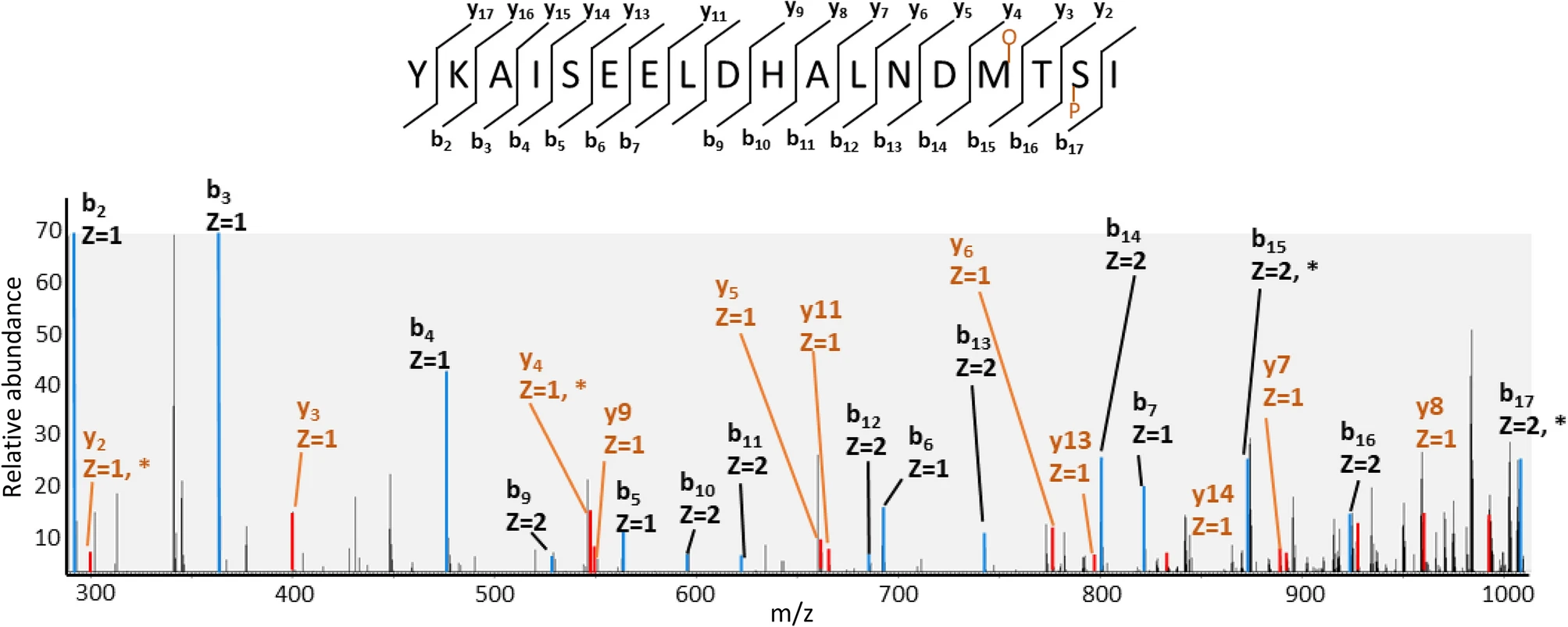
Bagwan, N. et al. Sci Rep. 2021.
Figure3. Fragmentation Spectra of Peptide with two PTMs: Phosphorylation and Oxidation
At MtoZ Biolabs, we are committed to delivering high-quality, high-impact post-translational modifications (PTMs) service, helping researchers unravel the complexity of protein modifications with unparalleled precision. Get in touch with us today to discuss your PTM analysis needs!
How to order?







