Protein Characterization by Mass Spectrometry Service
- Low-complexity samples: Gel spots or bands, approximately 1-2 cm in length.
- Medium-complexity samples: Protein mixtures, including column-purified samples.
- Moderate-to-high complexity samples: Protein mixtures such as IP-derived samples, secreted proteomes, or subcellular organelles (e.g., mitochondria, endoplasmic reticulum, lysosomes).
- High-complexity samples: Whole cell lysates, tissues.
Protein characterization by mass spectrometry is a comprehensive analytical method based on mass spectrometry, used to determine protein molecular weight, amino acid sequence, post-translational modifications, and structural features. This technique allows researchers to address complex challenges in protein identification, quantification, modification detection, and structural analysis, providing a deeper understanding of protein roles in biological processes.
MtoZ Biolabs offers Protein Characterization by Mass Spectrometry Service using advanced mass spectrometry technologies to provide high-quality analysis. Our Protein Characterization by Mass Spectrometry Service support a wide range of applications, from basic research to clinical needs, delivering precise insights into protein structure, function, interactions, and variations across different biological conditions.
Services at MtoZ Biolabs
Mass spectrometry works by measuring the intensity of ion signals to analyze the mass, abundance, and structure of proteins, offering comprehensive data for identification, quantification, and post-translational modification analysis. When paired with liquid chromatography (LC), mass spectrometry efficiently separates proteins from complex samples and provides accurate data for both quantitative and structural analyses.
In the Protein Characterization by Mass Spectrometry Service, MtoZ Biolabs employs state-of-the-art techniques such as LC-MS/MS, MALDI-TOF MS, and hydrogen-deuterium exchange mass spectrometry (HDX-MS), providing the following key services:
1. Protein Identification and Sequence Analysis
Accurately determine the amino acid sequence of proteins and identify protein variants, supporting functional studies and biomarker discovery.
2. Post-translational Modification Analysis
Examine phosphorylation, glycosylation, and other modifications, revealing functional regulation and intracellular signaling pathways.
3. Protein Structural Characterization
Use HDX-MS to provide detailed information on protein secondary and tertiary structures, aiding in the analysis of their spatial conformation and interactions with other molecules.
4. Protein Quantification
Utilize label-based (e.g., TMT, iTRAQ) and label-free quantification methods to accurately measure protein abundance across different samples.
5. Protein Interaction Analysis
Through immunoprecipitation coupled with mass spectrometry (Co-IP-MS), elucidate protein-protein interactions and signaling networks.
6. Protein Complex Analysis
Analyze the composition and functional role of protein complexes in biological processes, advancing understanding of their functional properties.

Figure 1. Workflow of Protein Characterization by Mass Spectrometry
Service Advantages
1. Advanced Analysis Platform: MtoZ Biolabs established an advanced mass spectrometry platform, guaranteeing reliable, fast, and highly accurate protein characterization service.
2. Multiplex Analysis Capability: Simultaneously quantify multiple target proteins, enhancing experimental efficiency and meeting the demands of complex sample analyses.
3. Broad Sample Compatibility: Our platform accommodates various biological samples—from purified proteins to complex matrices like cell lysates, tissues, and bodily fluids—to meet diverse research needs.
4. Comprehensive Data Support: Provide detailed insights from protein identification to advanced structural analysis, post-translational modifications, and interactions.
5. Customized Service: Tailored to the specific research needs of our clients, we offer flexible experimental design and personalized data analysis to ensure the achievement of research goals to the fullest extent.
6. One-Time-Charge: Our pricing is transparent, no hidden fees or additional costs.
Sample Submission Suggestions
We recommend the following categories of samples for optimal results in Protein Characterization by Mass Spectrometry Service:
For more specific sample submission requirements, please consult our technical team.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment
4. Data Analysis, Preprocessing, and Estimation
5. Bioinformatics Analysis
6. Raw Data Files
MtoZ Biolabs offers the Protein Characterization by Mass Spectrometry Service to comprehensively characterize protein properties, supporting both research and clinical applications. In addition to mass spectrometry analysis, we also offer other protein characterization methods, such as circular dichroism (CD) spectroscopy and nuclear magnetic resonance (NMR), helping you understand the properties and functions of proteins from different perspectives.
For any needs or inquiries, please feel free to contact us. MtoZ Biolabs is committed to providing professional technical support and high-quality services.
Case Study
Case 1: Application of Mass Spectrometry in Transgenic Crop Protein Characterization
This study utilized MALDI In-Source Decay MS technology to characterize proteins in transgenic crops, supporting the safety evaluation of genetically modified crops. By accurately analyzing the protein composition of crops, the study successfully identified key protein features that could impact food safety. This study demonstrated the value of the Protein Characterization by Mass Spectrometry Service in the safety evaluation of genetically modified crops and highlighted the important role of mass spectrometry in protein characterization and food safety detection.
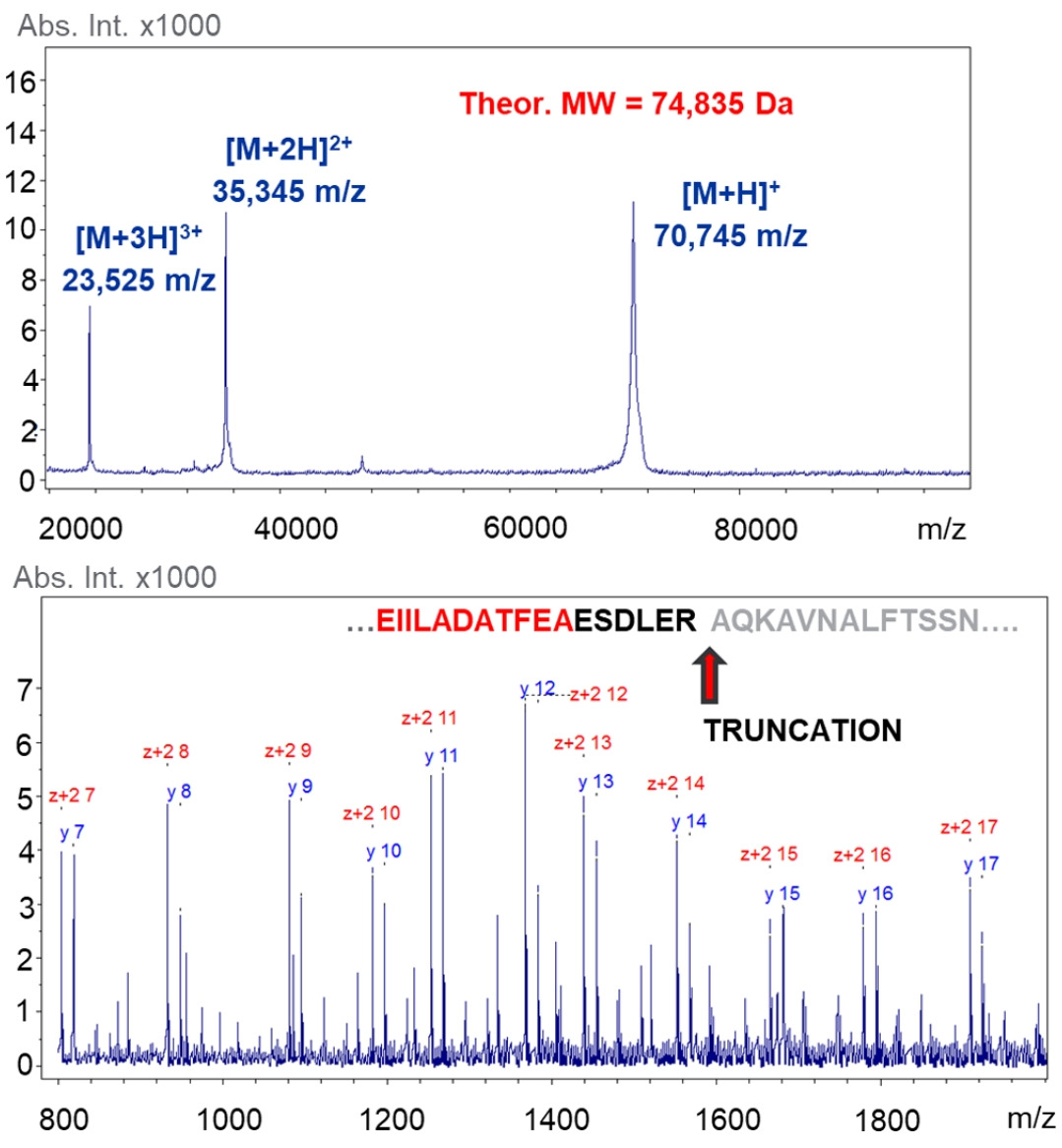
Birukou, I. et al. J. Agric. Food Chem. 2021.
Case 2: Application of Mass Spectrometry in Biopharmaceutical Protein Characterization
This study used isotope labeling (TMT) and high-resolution mass spectrometry to perform quantitative analysis of post-translational modifications in biopharmaceutical products. By leveraging the multiplex quantitative capabilities of TMT, the study achieved efficient quantification of protein modifications across multiple samples, providing precise data for biopharmaceutical quality control. This research demonstrated the application value of the Protein Characterization by Mass Spectrometry Service in the biopharmaceutical industry, particularly in drug development, quality assessment, and optimization.
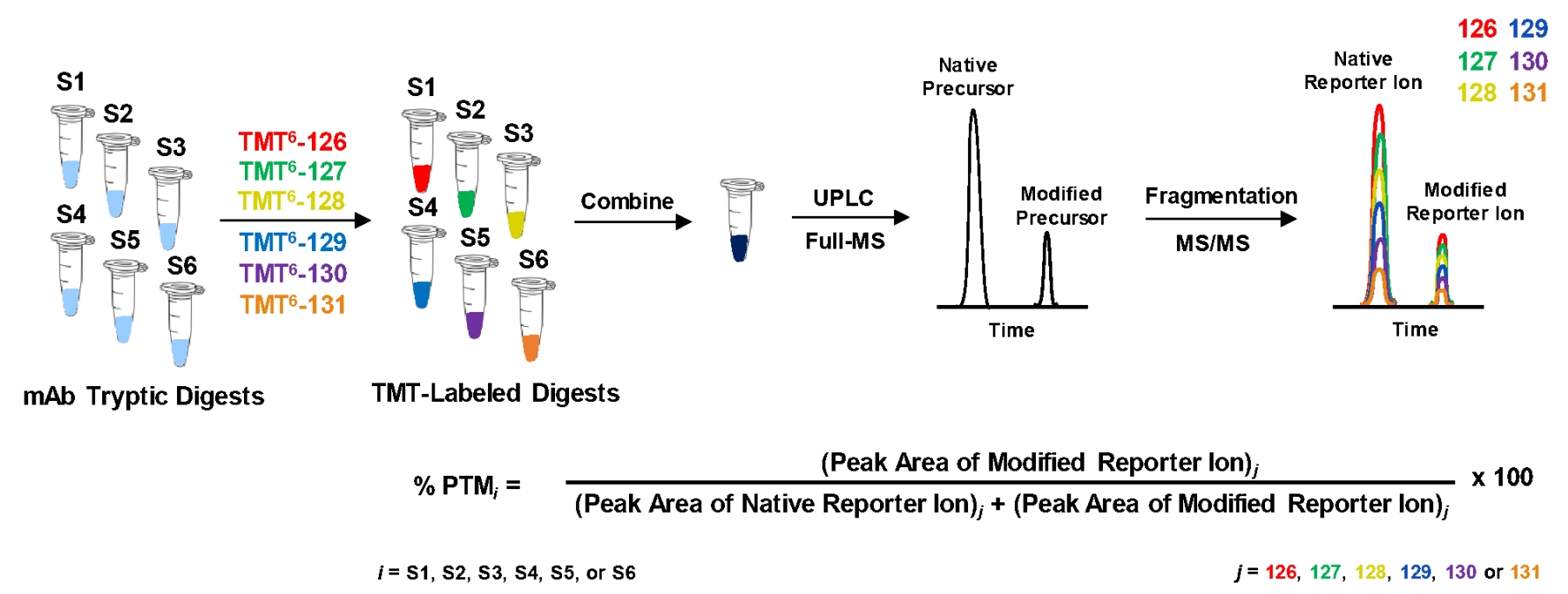
Mao, Y. et al. Anal. Chem. 2020.
FAQ
Q1: How is protein post-translational modification analyzed through mass spectrometry?
Our Protein Characterization by Mass Spectrometry Service use the Thermo Fisher Orbitrap Fusion Lumos mass spectrometer, combined with specific antibody enrichment techniques, to precisely analyze post-translational modifications of proteins. Through the enrichment of modifications like phosphorylation and glycosylation, we can quantitatively detect the modification types and their locations, revealing their regulatory effects on protein function.
How to order?







