SILAC Analysis Service
Stable isotope labeling with amino acids in cell culture (SILAC) is a technique where selected stable isotope-labeled amino acids are directly added to the cell culture medium. Compared to chemical labeling and label-free quantification techniques, it offers advantages such as simplicity, accurate quantification, and good reproducibility. Although mass spectrometry-based quantitative proteomics techniques have matured, the various physicochemical properties of peptides result in different spectral responses within the same mass spectrometer, making it difficult to analyze the relative abundance between different experimental samples. Stable isotope labeling in cell culture effectively addresses this issue by introducing stable isotopes into proteins or peptides, enabling relative quantification of peptides from different samples in the same mass spectrometry analysis. Currently, SILAC analysis service has been widely used in research fields such as characterizing proteomic changes between different biological samples, studying dynamic changes in protein post-translational modifications, and analyzing interacting proteins.
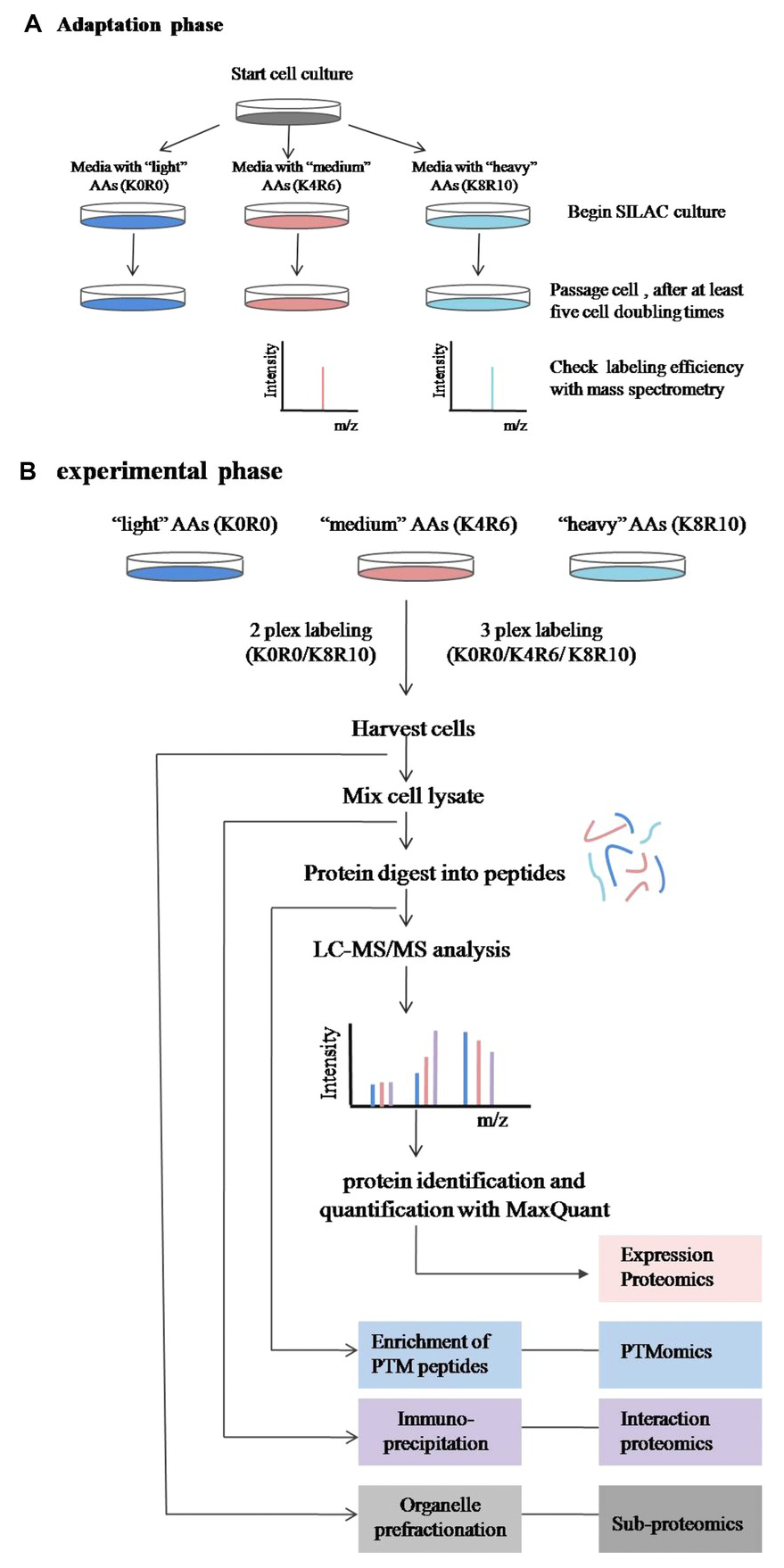
Chen, X. et al. Proteomics. 2015.
Figure 1. Experimental process of SILAC technology.
Experimental Principle
SILAC is the metabolic penetration of stable isotope labeled amino acids (such as 13C or 15N labeled arginine or lysine) into the entire proteome during protein metabolism, especially during cell culture. In the experiment, two cell populations are grown in two different media: the "light" medium contains naturally occurring amino acids, while the "heavy" medium contains stable isotope-labeled amino acids. After 5–6 generations of subculture, the cells grown in the heavy medium theoretically have all their proteins labeled with the heavy amino acids. After labeling, equal amounts of labeled and unlabeled cells or protein extracts are mixed, digested into peptides, and subjected to mass spectrometry. The quantitative principle of the SILAC analysis service is based on the ratio of labeled peptides to unlabeled peptides. Therefore, the relative abundance of peptides in the mixture can be quantified by comparing the signal intensities between the light and heavy samples. Since the number of cell divisions required for complete labeling depends on protein synthesis, degradation, and metabolism rates, labeling efficiency should be carefully tested before quantification.
Services at MtoZ Biolabs
Based on mass spectrometry technology combined with multi-dimensional data analysis, MtoZ Biolabs provides SILAC analysis service that allow for both protein identification and relative quantification through mass spectrometry, enabling the detection of differentially expressed proteins. However, this technique is only applicable to cell samples and requires prolonged cell culture. We are confident in providing professional and reliable services.
Service Advantages
1. High Labeling Efficiency
Living cell level labeling technology, stable labeling effect, labeling efficiency up to 100%.
2. High Sensitivity
The sample requirement is low, typically requiring only tens of micrograms of protein per sample.
3. High Throughput
Using mass spectrometry-based quantification, hundreds to thousands of proteins can be simultaneously identified and quantified.
4. High Precision
Multiple samples are mixed and processed simultaneously for separation, digestion, and identification, reducing the effects of experimental operations and instrument equipment, resulting in higher precision and reproducibility.
5. High Activity
The labeling employs in vivo techniques, providing a more accurate reflection of the sample's true physiological state.
Applications
1. Expression Proteomics
SILAC technology can monitor quantitative differences in protein levels under various conditions. It can also be used to identify differentially expressed proteins in organelles.
2. Dynamic changes in PTM
SILAC can be used for global and dynamic analysis of PTMs, including phosphorylation, acetylation, glycosylation, ubiquitination, and methylation.
3. PPI Research
SILAC analysis service can effectively differentiate between specific interacting proteins and non-specific proteins.
Case Study
1. SILAC-based Phosphoproteomics Reveals KSR1 Regulates DBC1’s Inhibitory Role in p53 Transcriptional Activity
KSR1 (a kinase suppressor of ras-1) is a potential regulatory gene in breast cancer. Quantitative mass spectrometry using SILAC technology to identify KSR1-regulated phosphoproteins in breast cancer. In vitro luciferase assays, co-immunoprecipitation, and western blotting were performed to further investigate KSR1’s role in breast cancer. Proteomics analysis showed that overexpression of KSR1 reduced phosphorylation at DBC1 (Deleted in Breast Cancer 1). Furthermore, KSR1 lowered p53 transcriptional activity by reducing phosphorylation at DBC1, leading to decreased interaction between DBC1 and Sirtuin-1 (SIRT1), which results in SIRT1-mediated deacetylation of p53. This case demonstrates how SILAC analysis service was used for phosphoproteomic analysis, revealing that KSR1 participates in regulating the p53 metabolic pathway in MCF7 cells.
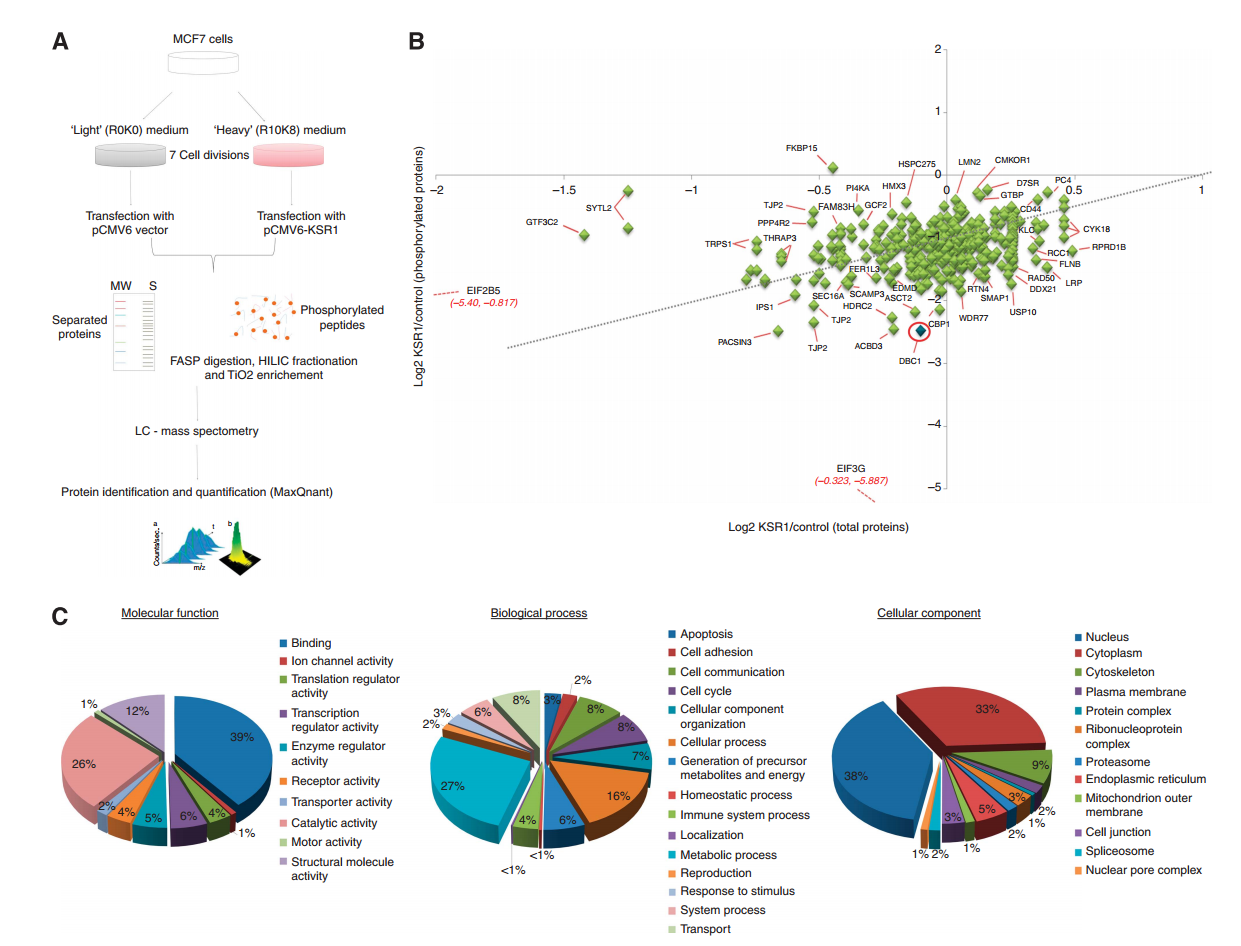
Zhang, H. et al. British Journal of Cancer, 2013.
Figure 2. Identification of KSR1-regulated phosphoproteome in breast cancer cells.
Deliverables
1. Labeling Test Results
2. Protein Identification and Quantification Results
3. Peptide Identification and Quantification Results
4. Differential Protein Results
5. Bioinformatics Analysis
6. Raw Data Files
7. Experimental Report
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details!
How to order?







