Advantages and Limitations of De Novo Protein Sequencing
De novo protein sequencing is an advanced technique for determining protein sequences without requiring genomic data or database references. This approach is particularly valuable for studying unknown proteins, novel antibodies, non-model organisms, and complex post-translational modifications (PTMs). Unlike traditional mass spectrometry-based identification, which relies on database searches, de novo sequencing is especially useful for novel protein discovery and modification analysis. However, challenges related to sequence assembly, detection of low-abundance proteins, interference from PTMs, and high costs hinder its broader application. This review systematically examines the strengths and limitations of de novo sequencing, providing insights into its applicability and future prospects.
Advantages of De Novo Protein Sequencing
1. Independence from Genomic and Database Constraints
Unlike traditional protein identification methods that require database matching, de novo sequencing directly determines protein sequences, making it indispensable for studying species with incomplete or missing genomic information.
2. High-Resolution Characterization of Post-Translational Modifications (PTMs)
(1) Open Modification Profiling: De novo sequencing does not require prior knowledge of PTMs. Instead, it identifies modifications by detecting characteristic mass shifts (e.g., phosphorylation +79.966 Da). This approach has enabled the discovery of previously unreported histone acetylation sites in tumor samples.
(2) Comprehensive Analysis of Multiple Modifications: It allows simultaneous characterization of coexisting PTMs on a single peptide (e.g., glycosylation and phosphorylation), overcoming the combinatorial challenges faced by traditional methods.
3. Antibody and Recombinant Protein Analysis
(1) Direct Sequencing of Variable Regions in Monoclonal Antibodies: De novo sequencing directly determines the complementarity-determining regions (CDRs) of hybridoma-derived or phage-displayed antibodies, eliminating the need for labor-intensive gene cloning.
(2) Quality Control of Recombinant Proteins: This method rapidly identifies sequence discrepancies caused by codon bias in expression systems, ensuring batch-to-batch consistency in biopharmaceutical production.
4. Applicability to Highly Complex Samples
(1) Proteomic Analysis of Low-Homology Species: De novo sequencing facilitates proteomic studies in extreme-environment organisms, such as deep-sea microbes or extraterrestrial samples, where genomic data are unavailable.
(2) Host-Pathogen Interaction Studies: It enables direct differentiation of host- and pathogen-derived proteins in viral infection research.
Limitations of De Novo Protein Sequencing
1. Challenges in Sequence Accuracy
De novo sequencing relies on MS/MS fragmentation patterns, but certain amino acids, such as leucine and isoleucine, have identical masses, making them indistinguishable by mass spectrometry alone. Additionally, some fragmentation methods may result in the absence of critical ion signals, compromising sequence assembly accuracy.
2. High Computational Complexity of Data Analysis
Unlike database-dependent identification, de novo sequencing requires complete sequence reconstruction, imposing significant computational demands. The complexity of mass spectrometry data, coupled with noise interference, necessitates the use of high-performance computing and advanced algorithm optimization to improve sequencing accuracy and efficiency.
3. Detection Limitations for Low-Abundance Proteins
Biological samples contain proteins spanning a wide dynamic range of abundance, with low-abundance proteins often being overshadowed by signals from highly abundant proteins. This signal suppression effect reduces sequencing sensitivity, posing challenges in detecting rare proteins and potential biomarkers.
4. Influence of Post-Translational Modifications on Fragmentation Patterns
PTMs can alter peptide fragmentation behavior, leading to incomplete b/y ion series. For instance, phosphorylation can cause selective suppression of specific fragment ions, complicating sequence interpretation and reducing overall sequencing accuracy.
5. High Costs and Extended Experimental Timelines
The operational costs of high-resolution mass spectrometers are substantial. Additionally, de novo sequencing involves multiple experimental steps, including enzymatic digestion, chromatographic separation, and extensive data processing, leading to prolonged experimental timelines. The computational resources required for sequence assembly further increase the financial burden of this approach.
Balancing Advantages and Limitations Across Different Applications
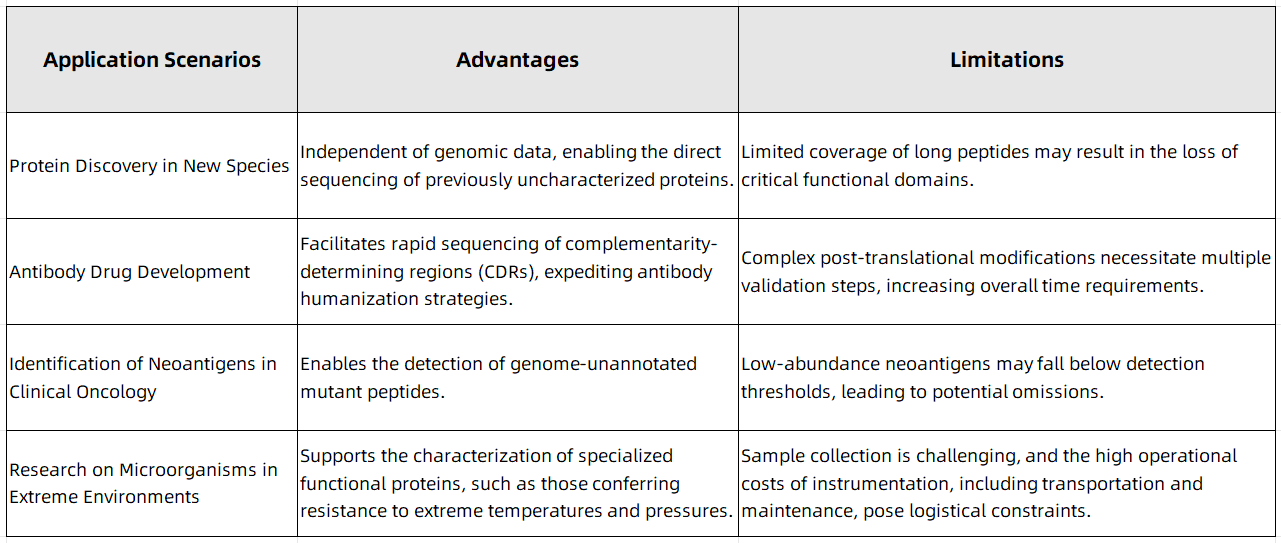
Figure 1
With continuous advancements in mass spectrometry hardware, breakthroughs in single-molecule sequencing technologies, and the increasing integration of artificial intelligence, de novo protein sequencing is anticipated to evolve from a specialized analytical technique into a comprehensive research platform. Over the next decade, its applications are expected to expand significantly in fields such as precision medicine and synthetic biology. MtoZ Biolabs offers high-precision and rapid de novo protein sequencing services, providing researchers with robust analytical solutions for comprehensive protein characterization.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







