LC MS Protein Identification Service
LC MS protein identification service combines liquid chromatography (LC) with mass spectrometry (MS) to efficiently separate and detect proteins and their fragments. LC MS is a cornerstone of modern proteomics, enabling the identification of protein components in complex biological samples. In this service, proteins in the sample are enzymatically digested into peptides, which are first separated by LC based on their chemical properties (e.g., hydrophilicity and molecular size). The separated peptides are then analyzed by a mass spectrometer, which detects their mass-to-charge ratio (m/z) and generates peptide mass spectra. Using these spectral data, the types, sequences and post-translational modifications of proteins can be identified. Leveraging chromatography and mass spectrometry platforms, MtoZ Biolabs offers LC MS Protein Identification Service providing clients with detailed insights into the composition, abundance, and function of proteins in samples.
Analysis Workflow
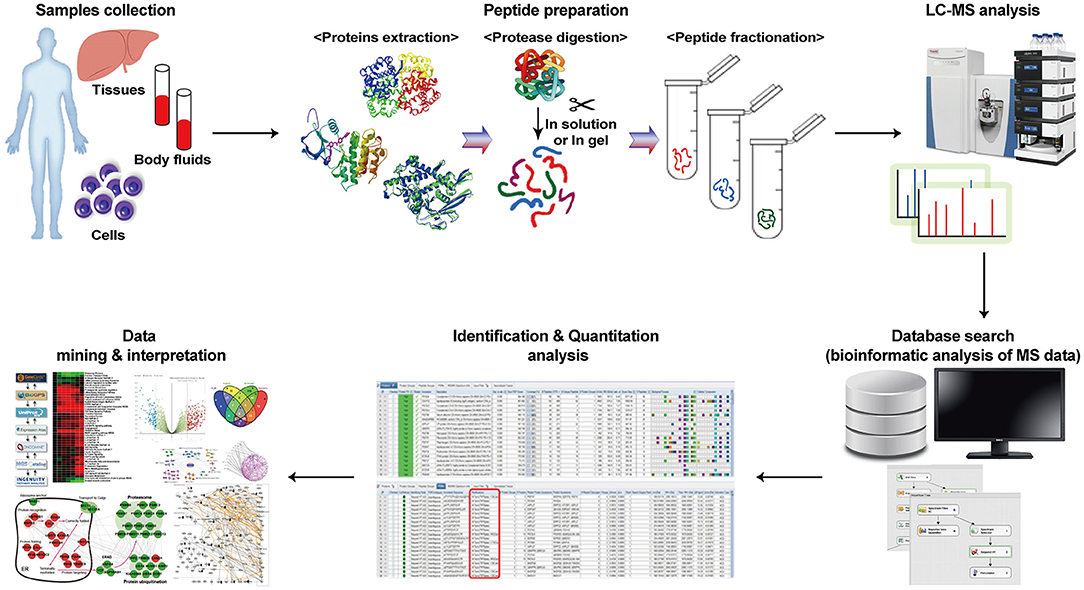
Kwon Y W. et al. Frontiers in medicine. 2021.
Sample Preparation
The sample undergoes extraction and purification, followed by chemical or enzymatic digestion to break down proteins into peptides.
Liquid Chromatography Separation
The peptides are separated using a liquid chromatography system based on their affinity differences, ensuring effective separation of distinct peptides.
Mass Spectrometry Analysis
The separated peptides are introduced into the mass spectrometer, where peptide mass spectra are generated through mass spectrometry analysis.
Data Analysis
High-efficiency bioinformatics tools are utilized to process the mass spectrometry data for protein identification and quantitative analysis.
Services at MtoZ Biolabs
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. MtoZ Biolabs can provide efficient and accurate LC MS protein identification service for proteins in samples such as protein extracts, SDS-PAGE protein bands, 2D protein spots, pull-down and Co-IP.
Applications
Common applications of LC MS protein identification service are as follows:
High-Throughput Protein Identification
Efficiently and accurately identifies proteins from complex biological samples.
Differential Protein Analysis
Analyzes differences in protein expression across samples, providing high-precision data for disease research and biomarker discovery.
Proteomics
Offers comprehensive analysis of protein composition within samples, revealing protein expression profiles.
FAQ
Q. How to Process Complex Biological Samples (e.g., Tissue Homogenates, Serum) to Reduce Background Interference and Enrich Target Proteins?
1. Sample Pretreatment
Desalting and Contaminant Removal: Use ultrafiltration, desalting columns, or protein precipitation methods to remove small-molecule impurities such as salts and metabolites, reducing background noise in mass spectrometry.
High-Abundance Protein Removal: For serum and plasma samples, apply immunoaffinity columns or Protein A/G enrichment techniques to deplete high-abundance proteins such as albumin and IgG.
2. Protein Enrichment and Separation
Subcellular Fractionation: Separate cellular components like nuclei, mitochondria, and endoplasmic reticulum using differential or gradient centrifugation to enrich specific proteins of interest.
Affinity Purification: Enrich specific proteins using antibodies or ligands. For example, phosphorylated proteins can be enriched with TiO₂ or IMAC columns.
SDS-PAGE or Gel Cutting: Combine electrophoresis with gel slicing techniques to reduce sample complexity and isolate target protein bands.
3. Optimize Protein Digestion
Ensure High Efficiency and Specificity: Use trypsin as the primary enzyme, and consider additional specific enzymes based on the characteristics of the target protein.
Prevent Non-Specific Degradation: Maintain low temperatures and use protease inhibitors to protect sample integrity and prevent unwanted protein breakdown.
By implementing these strategies, background interference can be effectively minimized, target proteins enriched, and mass spectrometry sensitivity and identification depth improved.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment (project-dependent)
4. Data Analysis, Preprocessing, and Estimation (project-dependent)
5. Bioinformatics Analysis
6. Raw Data Files
Case Study
This study utilized LC MS protein identification technology to conduct a comprehensive proteomics analysis of the host response in COVID-19 patients. The research identified key protein expression changes induced by viral infection, with a focus on uncovering critical pathways related to immune-inflammatory responses, coagulation dysfunction, and metabolic regulation abnormalities.
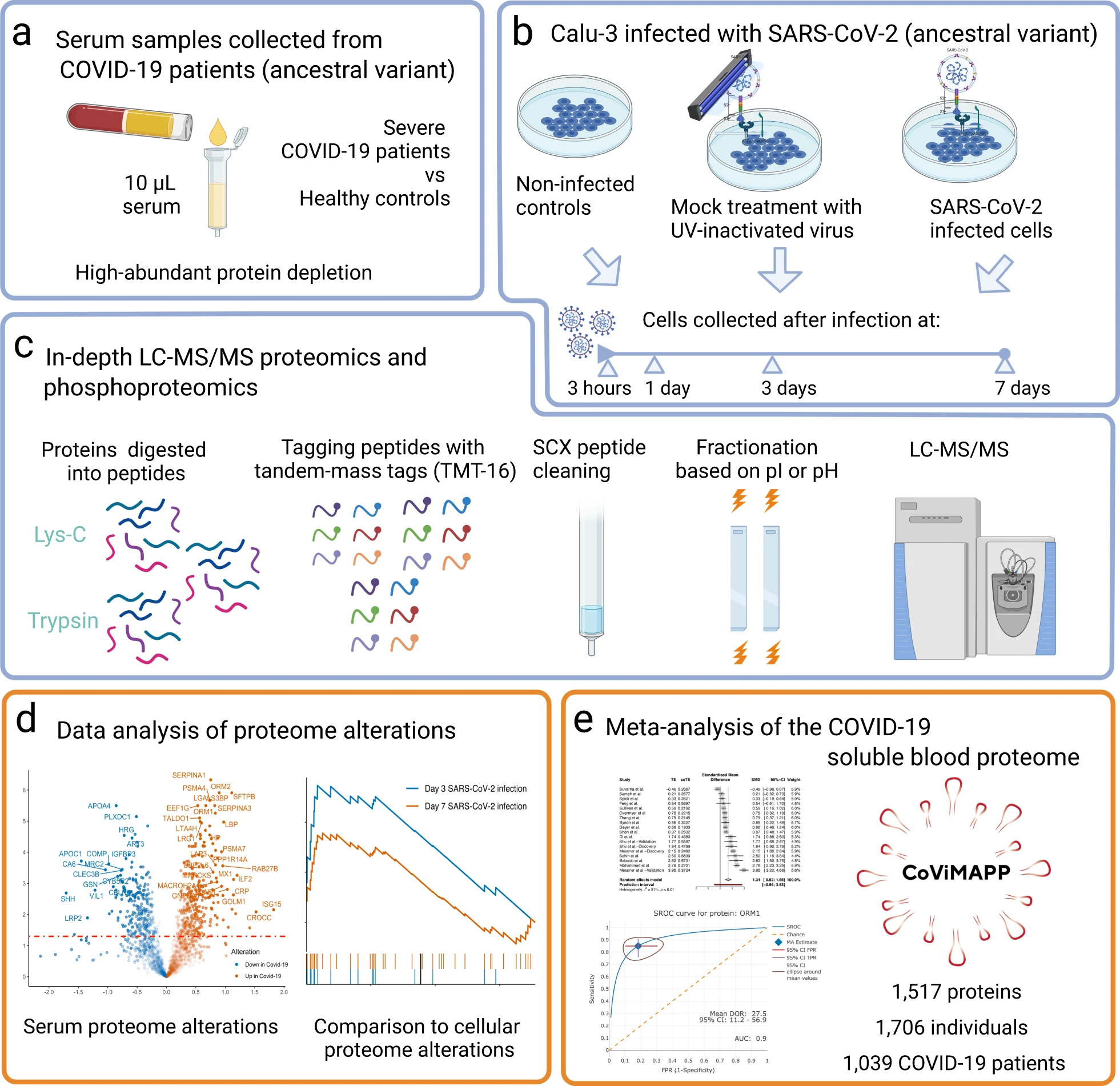
Babačić H. et al. Nat Commun. 2023.
How to order?







