N-Terminal Sequencing: A Comparative Analysis of Edman Degradation and Mass Spectrometry Approaches
N-terminal sequencing is a widely employed technique for determining the amino acid sequence of proteins. It plays a crucial role in proteomics, molecular biology, and biopharmaceutical research by elucidating protein structural characteristics, modifications, and post-translational processing. The primary methods used for N-terminal sequencing include Edman degradation, a classical chemical approach, and mass spectrometry, a modern analytical technique. These two methods differ markedly in their underlying principles, range of applications, and data interpretation strategies.
Edman Degradation
1. Principle
Edman degradation employs a chemical reaction in which phenyl isothiocyanate (PITC) reacts with the N-terminal amino acid residue, forming a cyclic intermediate. This intermediate is subsequently cleaved under acidic conditions, releasing a phenylthiohydantoin (PTH)-amino acid derivative, which is identified through high-performance liquid chromatography (HPLC).
2. Advantages
(1) Offers high sequence specificity and directly provides amino acid sequence information.
(2) Effective for sequencing purified protein samples without reliance on database searches.
(3) Enables analysis of post-translational modifications (PTMs) such as N-terminal acetylation and methylation.
3. Limitations
(1) Requires highly purified protein samples and is limited to sequencing peptides of 30–50 residues in length.
(2) Cannot be applied to N-terminally blocked proteins, including acetylated or cyclic peptides.
(3) Has a relatively long analytical process, making it less suitable for high-throughput applications.
Mass Spectrometry
1. Principle
Mass spectrometry-based N-terminal sequencing utilizes matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) or liquid chromatography-tandem mass spectrometry (LC-MS/MS) to analyze protein samples. By examining the mass-to-charge ratio (m/z) of peptide fragments generated through enzymatic digestion, and correlating these with database searches, the N-terminal sequence can be inferred.
2. Advantages
(1) Provides high sensitivity, enabling the detection of target proteins within complex mixtures.
(2) Well-suited for high-throughput analysis, allowing simultaneous sequencing of multiple proteins.
(3) Capable of identifying N-terminal modifications, including acetylation and glycosylation.
3. Limitations
(1) Heavily dependent on database searches, and does not yield direct primary sequence information.
(2) Susceptible to incomplete sequencing due to inefficient enzymatic digestion or limitations in mass spectrometry resolution.
(3) Requires sophisticated mass spectrometry instrumentation and advanced bioinformatics expertise for accurate data interpretation.
Comparative Analysis of Edman Degradation and Mass Spectrometry Approaches
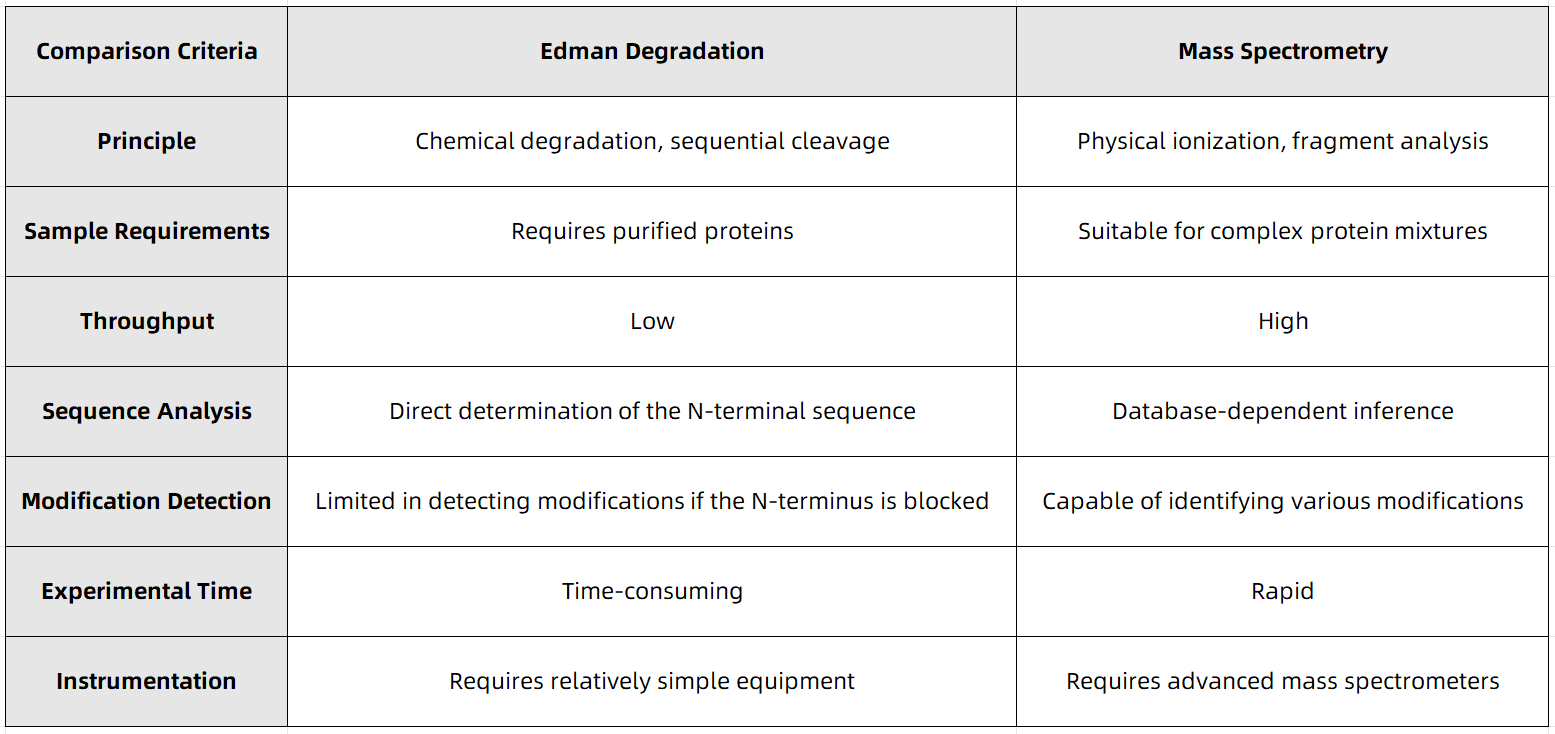
Figure 1
Both Edman degradation and mass spectrometry-based N-terminal sequencing exhibit distinct advantages and limitations, making them suitable for different experimental applications. Due to its high specificity and ability to determine the sequence directly, Edman degradation remains a valuable tool for analyzing N-terminally blocked proteins and small-scale protein studies. In contrast, mass spectrometry, characterized by its high throughput and sensitivity, has become the dominant method in modern proteomics research.
Researchers should select the most suitable strategy based on the specific requirements of their experiments to obtain accurate protein sequence information. MtoZ Biolabs provides high-precision N-terminal sequencing services with reliable and high-quality results. Contact us for further details!
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







