Native Mass Spectrometry: Principles, Advantages, and Applications
Native mass spectrometry is an emerging technique in mass spectrometry that allows for the direct analysis of intact protein molecules, providing insights into their native structures and post-translational modifications. It has become increasingly significant in proteomics research. In the following sections, we discuss its fundamental principles, advantages for proteomics, and various applications.
Principles
The core principle of mass spectrometry is to ionize the sample and measure its mass-to-charge ratio (m/z) using a mass spectrometer, thereby facilitating molecular identification and quantification.
1. Differences Between Traditional and Native Mass Spectrometry
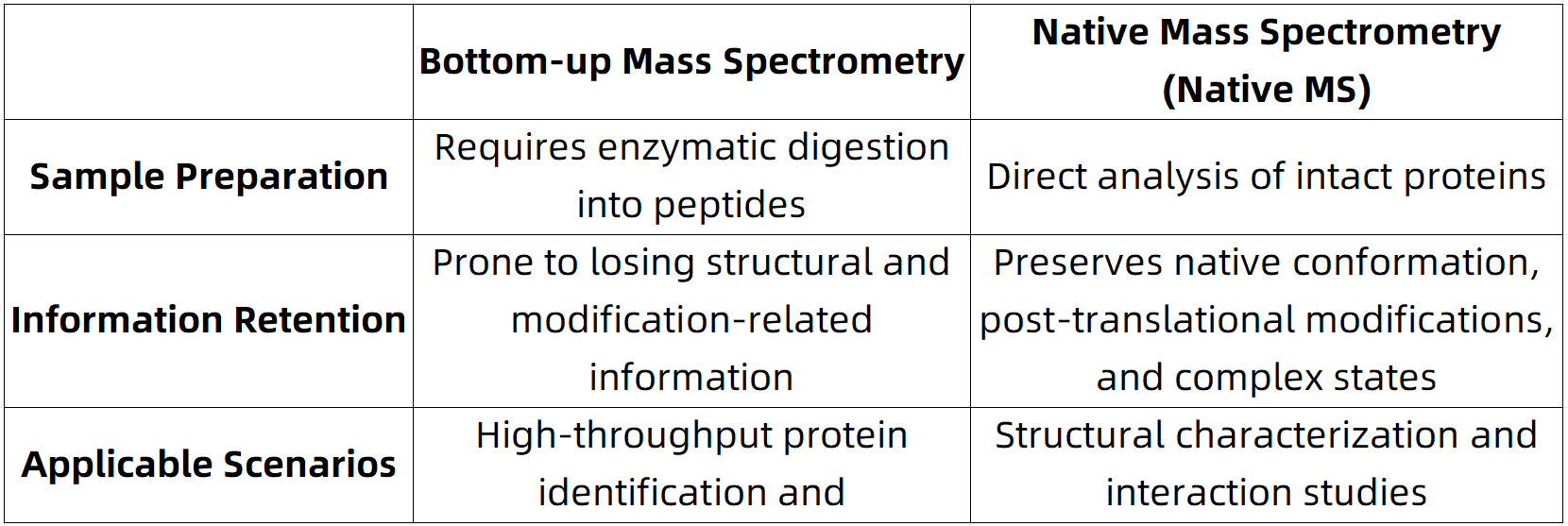
Figure 1
2. Technical Approaches
(1) Low-Energy Ionization: Gentle ionization techniques, such as electrospray ionization (ESI), are employed to preserve non-covalent interactions in proteins (e.g., hydrogen bonds and hydrophobic interactions).
(2) High-Resolution Mass Spectrometers: High-resolution instruments, such as Orbitrap and Fourier Transform Ion Cyclotron Resonance (FT-ICR) mass spectrometers, enable precise macromolecular mass determination, reaching the megadalton (MDa) range.
Advantages
1. Preservation of Native Protein Structure and Modifications
Traditional mass spectrometry based on enzymatic digestion relies on fragmenting proteins into small peptides, often resulting in the loss of modification information. In contrast, native mass spectrometry directly analyzes intact proteins without enzymatic digestion, thereby maintaining their native three-dimensional structure and post-translational modifications more effectively. For modifications such as phosphorylation, acetylation, and glycosylation, Native MS enables more comprehensive and accurate detection and quantification.
2. Enhanced Capability for Complex Sample Analysis
In complex biological samples, traditional enzymatic digestion methods may fail to detect certain proteins due to their low abundance or incomplete modifications. By eliminating the digestion step, native mass spectrometry allows for the direct detection and analysis of complex proteomes, particularly those containing multiple modifications or exhibiting high heterogeneity. This approach enables a more comprehensive characterization of proteins, particularly full-length proteins.
3. Analysis of Protein-Protein Interactions
Protein-protein interactions play a fundamental role in cellular functions and biological processes. Native mass spectrometry preserves the non-covalent binding states of protein interactions, facilitating the study of their underlying mechanisms. Through mass spectrometric analysis, researchers can simultaneously detect multiple protein components within a complex, providing a powerful tool for mapping protein-protein interaction networks.
4. High-Resolution Analysis
Native mass spectrometry typically requires high-resolution mass spectrometers and precise analytical techniques. High-resolution mass spectrometric data enable researchers to obtain detailed molecular information on proteins, particularly when analyzing large macromolecular proteins and complex modifications, allowing for the effective differentiation of subtle structural variations. This capability positions Native MS as a valuable tool in large-scale proteomics research and precision medicine applications.
Applications
1. Analysis of Post-Translational Modifications
Post-translational modifications (PTMs) regulate protein function through various chemical modifications, including phosphorylation, glycosylation, methylation, and acetylation. Conventional enzymatic digestion-based mass spectrometry may fail to comprehensively capture complex PTM patterns. In contrast, native mass spectrometry, which does not require enzymatic digestion, preserves PTM information more effectively. This advantage is particularly evident in the detection of phosphorylation and acetylation, where native MS offers superior accuracy and reliability. These capabilities enhance our understanding of intracellular signaling pathways and metabolic regulation.
2. Investigation of Protein-Protein Interactions
Protein-protein interactions are fundamental to cellular function, signal transduction, and disease progression. Traditional enzymatic digestion-based mass spectrometry often encounters limitations in accurately characterizing protein complexes and their binding interactions. Native mass spectrometry enables direct analysis of intact protein complexes, providing critical insights into protein-protein interactions, particularly non-covalent associations. This approach facilitates the elucidation of protein interaction networks in biological systems, aids in biomarker discovery, and contributes to the identification of novel drug targets.
3. Discovery of Biomarkers for Cancer and Other Diseases
Proteomics plays a crucial role in early disease diagnosis and therapeutic strategy development. Native mass spectrometry enables the identification of disease-associated PTMs and structural alterations in intact proteins, many of which serve as early indicators of pathological processes. In oncology research, native MS provides a powerful approach to detect PTM dysregulation and conformational changes in tumor-related proteins, aiding in the discovery of novel biomarkers for early cancer detection and precision medicine applications.
4. Development and Characterization of Biopharmaceuticals
Biopharmaceuticals, including monoclonal antibodies and vaccines, have seen significant advancements in recent years. Native mass spectrometry offers a powerful analytical tool for characterizing the structural integrity, PTM profiles, and stability of these biologics. In drug development, this technique plays a vital role in optimizing biopharmaceutical formulations, ensuring structural fidelity, and improving therapeutic efficacy, particularly during early-stage quality control and lead optimization processes.
By preserving and analyzing the native conformation and PTMs of proteins, native mass spectrometry provides precise insights into protein function, interactions, and modifications, as well as their roles in disease pathogenesis. MtoZ Biolabs, leveraging high-resolution mass spectrometry platforms, offers advanced native MS-based protein characterization services to support scientific research and biopharmaceutical development.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







