SILAC Assay Service
Stable Isotope Labeling by Amino Acids in Cell Culture (SILAC) Analysis is a key technique in the field of proteomics. By integrating the principles of stable isotope labeling with refined cell culture strategies, SILAC enables an in-depth investigation of intracellular proteins. Simply put, this approach leverages amino acids labeled with stable isotopes, allowing cells to incorporate these uniquely marked amino acids into newly synthesized proteins during their natural growth and metabolism. This, in turn, facilitates precise and specific tracking, identification, and highly accurate quantification of these labeled proteins.
SILAC assay service focuses on protein synthesis dynamics, various modification patterns, and precise functional state changes, shedding light on how proteins exert their biological effects in the intricate cellular microenvironment. By combining isotope labeling with cell culture, this technique provides keen insights into subtle protein fluctuations and real-time dynamic changes under diverse conditions, such as cellular stress responses, specific physiological transitions, and pathological state evolution.
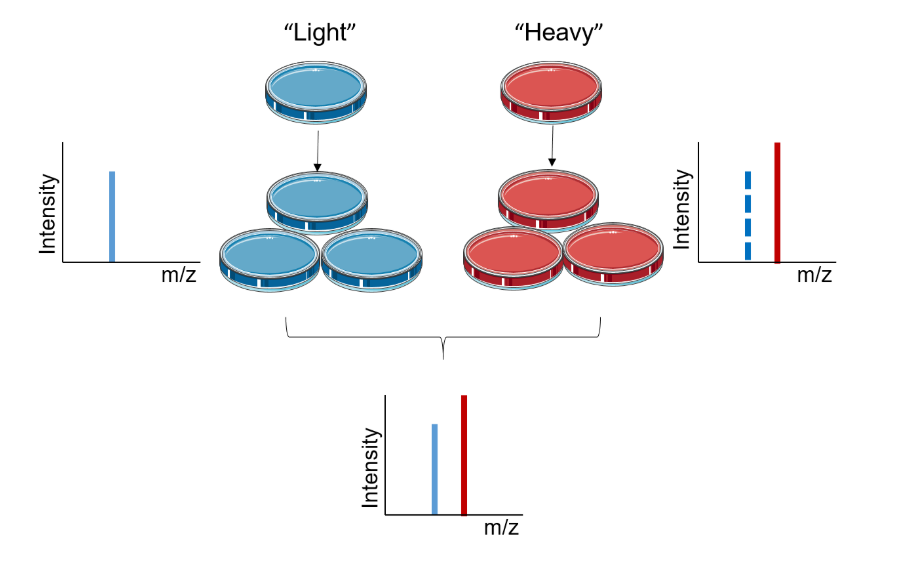
Chen X, et al. Proteomics. 2015.
Figure 1. Principles of SILAC Technology.
Services at MtoZ Biolabs
Utilizing Thermo’s Orbitrap series mass spectrometers in combination with a Nano-LC system, MtoZ Biolabs provides SILAC assay service that ensure high accuracy and sensitivity of data acquisition. Our key services include:
1. Proteomics Analysis Services
(1) Quantitative Analysis: SILAC assay service enables precise protein labeling, followed by mass spectrometry (MS) detection, where peptides carrying different isotopic labels are separately quantified. This allows for the calculation of relative or absolute protein abundance, facilitating the comparison of protein expression levels under different cellular states or experimental conditions.
(2) Post-Translational Modification (PTM) Analysis: SILAC assay service can be applied to study protein phosphorylation, methylation, acetylation, and other post-translational modifications (PTMs). By identifying and quantifying modification sites, researchers can gain insights into the role of PTMs in cellular signaling, metabolic regulation, and other biological processes.
2. Protein-Protein Interaction Analysis Services
(1) SILAC-Co-Immunoprecipitation Mass Spectrometry (SILAC-COIP-MS): The SILAC method is used to label both the experimental and control groups, followed by co-immunoprecipitation (CO-IP) to capture interacting proteins. Mass spectrometry is then employed for qualitative and quantitative analysis of these interactions. This approach is particularly useful for studying protein interaction networks and identifying proteins that specifically bind to a target protein.
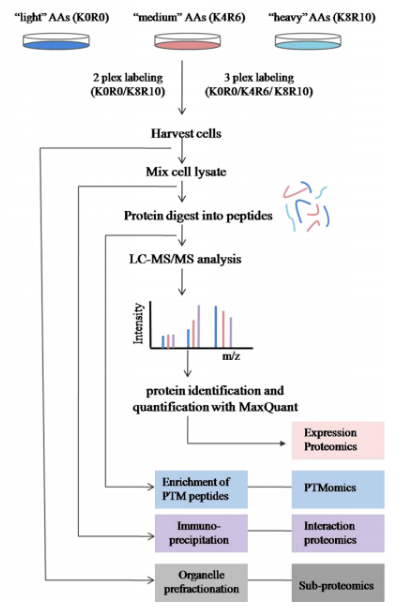
Chen, X. et al. Proteomics, 2015.
Figure 2. Workflow for SILAC Analysis.
Service Advantages
1. High Efficiency
MtoZ Biolabs' past projects demonstrate high labeling efficiency, unaffected by lysis buffer components, ensuring stable labeling performance.
2. High Accuracy
Quantitative accuracy and repetition are high, and subsequent experiments such as proteolytic are performed after mixing samples and other similarities, with small errors.
3. High Sensitivity
Requires minimal sample input, with cell quantities reaching as low as the single-cell level.
4. Post-Translational Modification (PTM) Detection
MtoZ Biolabs provides comprehensive PTM analysis, including phosphorylation, ubiquitination, and glycosylation, enabling a thorough investigation of protein modifications.
5. High Data Quality
Ensures deep data coverage with strict quality control. MtoZ Biolabs utilizes an AI-driven bioinformatics platform to integrate all high-throughput proteomics data, providing a comprehensive analytical report.
6. Customized Research Solutions
MtoZ Biolabs offers tailored solutions to address your specific research questions and experimental requirements.
Sample Submission Suggestions
1. Sample Type
We accommodate all types and quantities of cells. Ensure the samples are properly preserved to maintain protein integrity and spatial distribution.
2. Sample Amount
More than 3 biological replicates are recommended
Note: If you have any special requirements or need assistance with sample preparation, please feel free to contact us.
Applications
1. Expression Proteomics
SILAC provides an in vivo strategy for labeling proteins with different stable isotope forms of amino acids, allowing for quantitative monitoring of protein level variations under different conditions. Additionally, SILAC assay service is used to identify differentially expressed proteins within specific cellular compartments, such as the nucleus, nucleolus, or insulin secretion granules in cells.
2. Dynamic Changes in Post-Translational Modifications (PTMs)
For PTM analysis, SILAC-labeled peptides undergo fractionation and enrichment steps to enhance the identification of modified peptides. Combined with mass spectrometry (MS) technology, SILAC assay service enables comprehensive and dynamic analysis of PTMs, including phosphorylation, acetylation, glycosylation, ubiquitination, and methylation.
3. Protein-Protein Interaction Studies
In protein-protein interaction studies, protein complexes are immunoprecipitated from a mixture of SILAC-labeled cell lysates. When combined with SILAC, this approach effectively differentiates specific interaction partners from non-specific background proteins. The abundance of specific interacting proteins purified from the bait sample is significantly higher than in the control sample, resulting in a quantitative ratio far greater than 1. In contrast, non-specific background proteins should have comparable abundance in both bait and control samples, yielding a ratio close to 1. SILAC-based quantitative proteomics is applicable for studying exogenous, endogenous, or inducible interacting proteins.
Case Study
1. YiaC and CobB Regulate Lysine Lactylation in Escherichia coli
Lysine lactylation (Kla) has been previously reported to play a role in transcriptional regulation in human cells. however, its characteristics, regulatory mechanisms, and functional consequences in prokaryotes remain unclear. This study identified the regulatory enzyme system of Kla in Escherichia coli, demonstrating that YiaC and CobB function as a lysine lactylase and lysine delactylase, respectively, catalyzing the addition and removal of Kla both in vitro and in vivo. Furthermore, the study utilized SILAC technology to analyze the effects of YiaC or CobB gene deletion on lysine lactylation levels in E. coli. The results led to the construction of an enzyme-regulated Kla molecular network and revealed that CobB-mediated K382la modification of pyruvate kinase PykF plays a key role in regulating bacterial metabolism and growth.
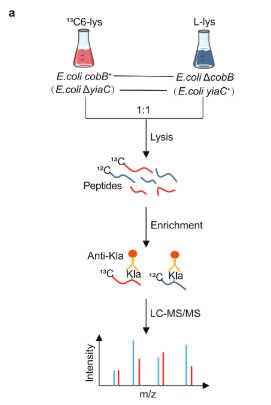
Dong, H. et al. Nat Commun, 2022.
Figure 3. Understanding the Specific Applications of SILAC Analysis.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment
4. Data Analysis, Preprocessing, and Estimation
5. Bioinformatics Analysis
6. Raw Data Files
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details!
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
Label-Free Quantitative Proteomics Service, MS Based
How to order?







