Resources
Proteomics Databases
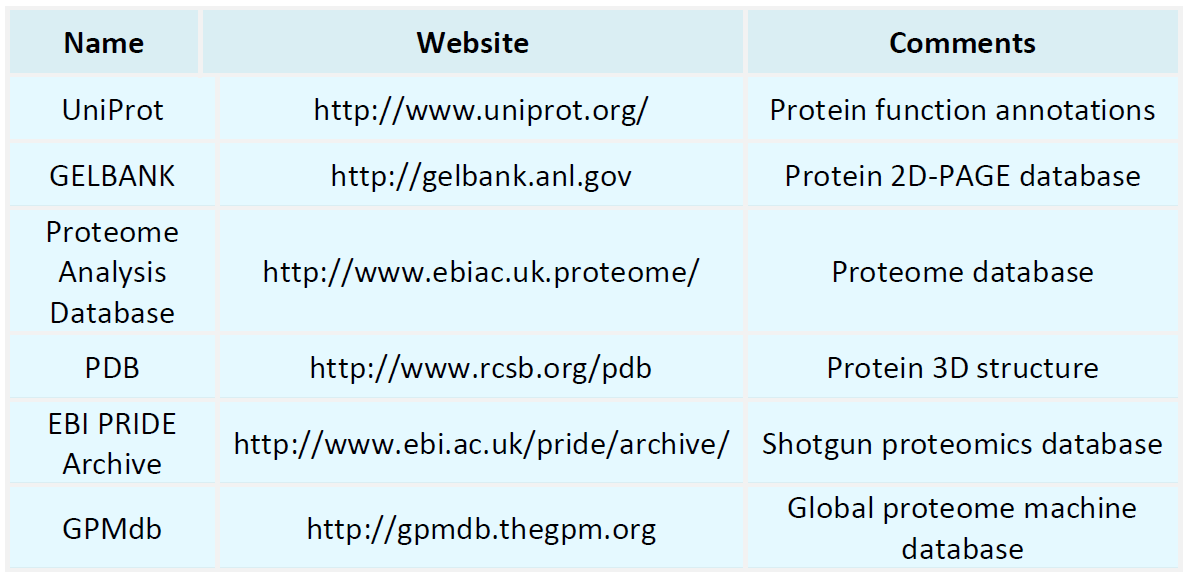
Metabolomics Databases
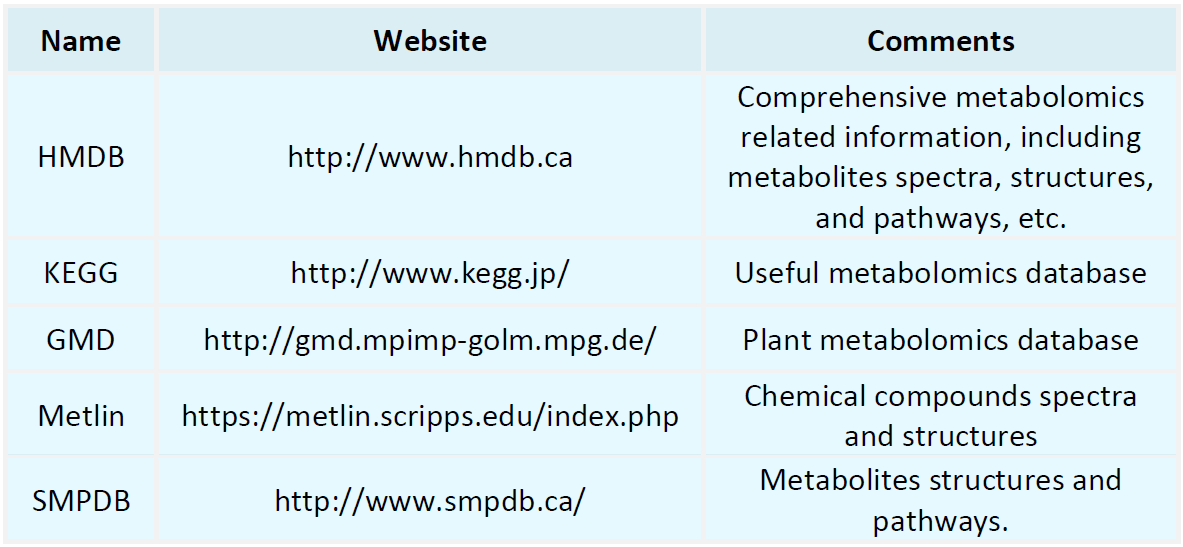
-
Protein Expression Studies are critical area of study in biology, focusing on the processes of synthesis, folding, modification, and degradation of proteins within organisms. Proteins are essential to life, facilitating cellular functions, signal transduction, and metabolic regulation. By studying protein expression, scientists can better understand genomic functions and elucidate how genetic mutations contribute to diseases. This research not only unveils fundamental life principles but also underpin......
-
• Pea Protein Complete Amino Acid Profile
Pea protein complete amino acid profile is a method used to determine the amino acid composition of pea protein. This process can reveal the nutritional value of pea protein in detail and provide valuable information about its functional properties in different applications. Pea protein is an important source of plant protein, attracting attention for its rich nutritional components and excellent digestibility. Through pea protein complete amino acid analysis, scientists and producers can obtain compr......
-
• Mass Spectrometry Peptide Fragmentation
Mass spectrometry peptide fragmentation is an essential technique in proteomics, primarily used to discern the primary structure of peptides. This method involves three main steps: peptide ionization, selection, and fragmentation. Initially, peptides are ionized to form charged ions, which are then selected for analysis using the mass spectrometer's mass selection capabilities. Subsequently, these ions are fragmented into smaller components via techniques such as collision-induced dissociation (CID), ......
-
• Membrane Proteomics Overview
Membrane proteomics is a branch of omics technology that studies all proteins in biological membranes. Biological membranes are structures composed of lipid bilayers and proteins, which exist in cell membranes and organelle membranes. Membrane proteins account for about 30% of all encoded proteins. However, due to their special biochemical properties, such as hydrophobicity and low solubility, they are often overlooked in studies. Membrane proteomics systematically identifies, quantifies, and investig......
-
• MALDI-TOF Microbial Identification
Maldi-Tof Microbial Identification is a pivotal technology, offering rapid and precise results. By analyzing the mass spectrometry profiles of microbial proteins, MALDI-TOF MS enables the differentiation of various microorganism species. Its applications span clinical microbiology, food safety, environmental monitoring, and biopharmaceuticals. In clinical settings, MALDI-TOF microbial identification swiftly identifies pathogens, providing clinicians with diagnostic insights to develop effective treatm......
-
Maldi tof spectroscopy is a rapidly advancing technology in bioanalysis. It offers unique advantages in areas such as proteomics, microbial identification, and drug metabolism research. MALDI-TOF MS allows for rapid and accurate mass analysis and component identification of samples. The technology, traced back to the 1980s, has expanded its applications with ongoing advances, becoming an essential tool in biomedical and chemical research. In proteomics, it is extensively used for protein identificatio......
-
• MALDI Microbial Identification
Matrix-Assisted Laser Desorption/Ionization (MALDI) is an analytical technique employed for the classification and identification of microorganisms. This method integrates laser desorption/ionization with time-of-flight mass spectrometry, utilizing matrix materials to absorb laser energy and transfer it to sample molecules, thereby achieving efficient ionization. The uniqueness of MALDI Microbial Identification lies in the use of matrix materials, which can effectively absorb laser energy and transfer......
-
• Maldi Tof Identification Microorganisms
Maldi Tof Identification Microorganisms is a method that uses Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS) to rapidly identify and classify microorganisms. The core principle is to ionize the proteins in the microbial sample using laser energy, and then analyze these ions through a time-of-flight mass spectrometer. Maldi Tof Identification Microorganisms has a wide range of applications in medical diagnostics, food safety, environmental monitoring, and bi......
-
2D proteomics is used to isolate and identify proteins in complex biological samples. The full name of the technique is Two-Dimensional Gel Electrophoresis (2-DE) . 2D proteomics combines two separation methods: isoelectric focusing and SDS-PAGE (polyacrylamide gel electrophoresis) to effectively separate and analyze protein molecules. The core value of this method lies in its ability to isolate thousands of proteins under a single experimental condition, allowing researchers to comprehensively analyz......
-
Terminal Sequencing is a method used for analyzing the amino acid sequence of protein or peptide molecules, with a particular focus on determining the N-terminal (amino end) and C-terminal (carboxyl end) amino acid sequences of these molecules. The main applications of terminal sequencing technology include protein identification and characterization, protein structure and function research, and biomarker discovery. In drug development, this technology can be used to determine the structure of target ......
How to order?







