Resources
Proteomics Databases
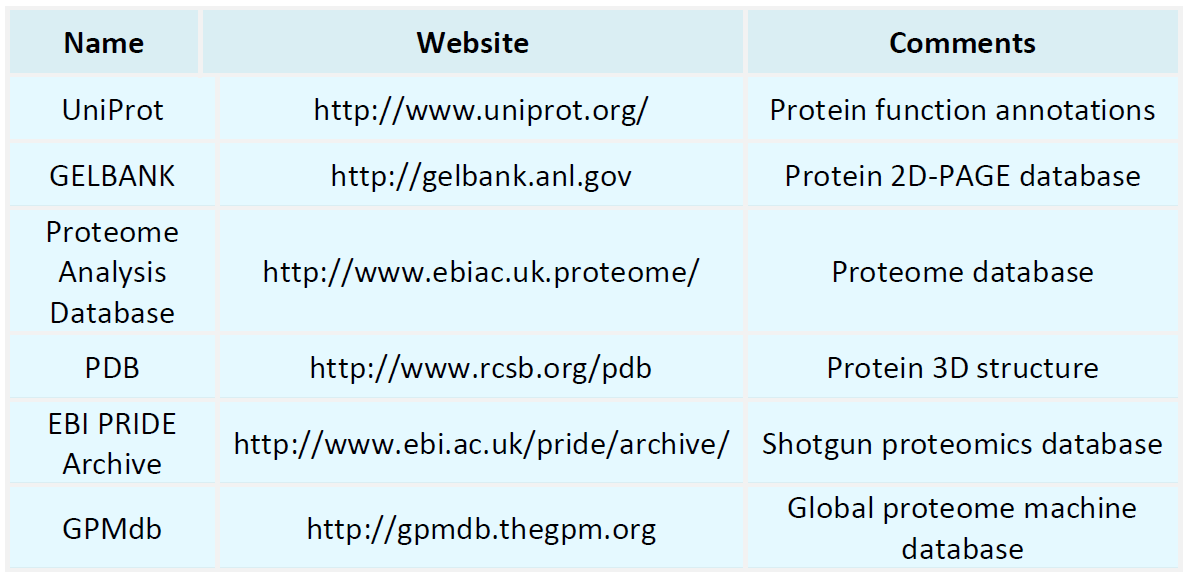
Metabolomics Databases
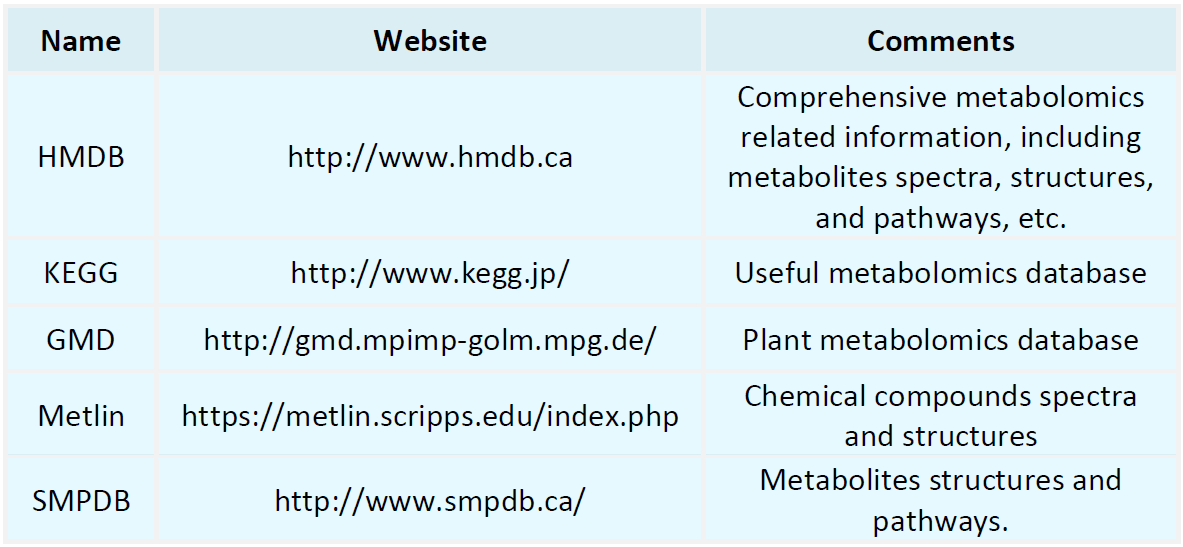
-
• How to Process Shotgun Proteomics Data in Multi-Omics Integration?
In multi-omics integration analyses, appropriate processing of Shotgun proteomics data is of critical importance. Proteomics data are inherently semi-quantitative, characterized by a high proportion of missing values and substantial variability across samples and analytical platforms. Inadequate data handling can therefore substantially impair integrative analyses with other omics layers, including genomics, transcriptomics, and metabolomics. Shotgun Proteomics: Data Characteristics and Challenges Sh......
-
• Disulfide Bonds in Proteins: Formation, Function, and Analytical Methods
Within the precise architecture of biomolecules, disulfide bonds are often overlooked yet fundamentally important chemical linkages. They are formed through covalent bonding between the sulfur atoms of two cysteine residues and can significantly enhance the structural stability and functional robustness of proteins. In particular, in key biomolecules such as secretory proteins, therapeutic antibodies, and biological enzymes, disulfide bonds frequently serve as molecular locks that ensure correct prote......
-
• Membrane Protein–Molecule Interaction Analysis
Membrane proteins account for nearly 30% of all proteins encoded by the human genome, and they constitute more than 60% of current drug targets. They are not only involved in essential physiological processes such as substance transport, signal transduction, and cell recognition, but also serve as key nodes in a wide range of diseases, including viral invasion, cancer metastasis, and neurodegenerative disorders. The interactions between membrane proteins and ligands, signaling molecules, and other pro......
-
• How to Achieve Quantitative Mapping of Protein Localization in Tissues?
Quantitative mapping of protein localization represents a critical foundation for understanding protein function, regulatory mechanisms, and cellular behavior under pathological conditions. With recent advances in mass spectrometry (MS) technologies, it has become feasible to perform high-throughput and quantitative analyses of protein localization at the subcellular level. Core Significance of Quantitative Mapping of Protein Localization Cellular function critically depends on accurate protein local......
-
• Comprehensive Guide to Protein Glycosylation Analysis (LC‑MS/MS Workflow)
Protein glycosylation is a fundamental post-translational modification (PTM) that is widely distributed in eukaryotic cells. It plays critical roles in regulating protein folding, stability, subcellular localization, and molecular interactions. Glycosylation has become increasingly important in biopharmaceutical development, cancer biomarker discovery, and vaccine research. Compared with other common PTMs such as phosphorylation and acetylation, glycosylation exhibits pronounced structural diversity, ......
-
• What Are Targeted Post-Translational Modifications?
Post-translational modifications (PTMs) are chemical alterations that occur to proteins after translation, including phosphorylation, acetylation, ubiquitination, and others. These modifications not only regulate protein activity, subcellular localization, and stability, but also exert substantial influence on signaling pathways, cellular differentiation, and disease processes. In proteomics, targeted post-translational modifications analysis refers to the selection of specific PTM classes and defined......
-
• How to Apply Targeted Mass Spectrometry for PTM Verification?
Protein post-translational modifications (PTMs) constitute fundamental regulatory mechanisms underlying essential biological processes, including cell fate determination, signal transduction, and metabolic homeostasis. Common PTMs include phosphorylation, acetylation, ubiquitination, and methylation. Although high-throughput mass spectrometry approaches, such as data-dependent acquisition (DDA) and data-independent acquisition (DIA), are widely used for the initial discovery of PTM sites, subsequent v......
-
• How to Identify Protein–Protein Interactions Using Mass Spectrometry?
Protein–protein interactions (PPIs) are central to biological processes such as cellular signal transduction, metabolic regulation, and chromatin remodeling. Identifying and characterizing protein interaction networks not only facilitates mechanistic understanding of cellular functions but also plays an important role in drug-target screening and studies of disease mechanisms. With the rapid advancement of mass spectrometry (MS) in proteomics, MS has become one of the most widely used and robust appro......
-
• What Is the Workflow for Targeted PTM Analysis?
Post-translational modifications (PTMs) participate extensively in fundamental biological processes, including cellular signal transduction, metabolic regulation, and gene expression. Their dynamic alterations often serve as important molecular indicators of disease onset, progression, and responses to therapy. With rapid advances in mass spectrometry (MS), targeted PTM analysis has become a powerful approach for elucidating regulatory mechanisms in complex biological systems. In contrast to global pr......
-
• How to Improve Sensitivity in Targeted PTM Detection by Mass Spectrometry?
Protein post-translational modifications (PTMs), including phosphorylation, acetylation, ubiquitination, and glycosylation, play pivotal roles in regulating signal transduction, cellular metabolism, and disease pathogenesis. Relative to unmodified proteins, PTM-bearing proteins/peptides are frequently low in abundance, exhibit heterogeneous modification sites, and may be chemically or enzymatically unstable; consequently, achieving high sensitivity and specificity remains a central challenge. Owing to......
How to order?







