Resources
Proteomics Databases
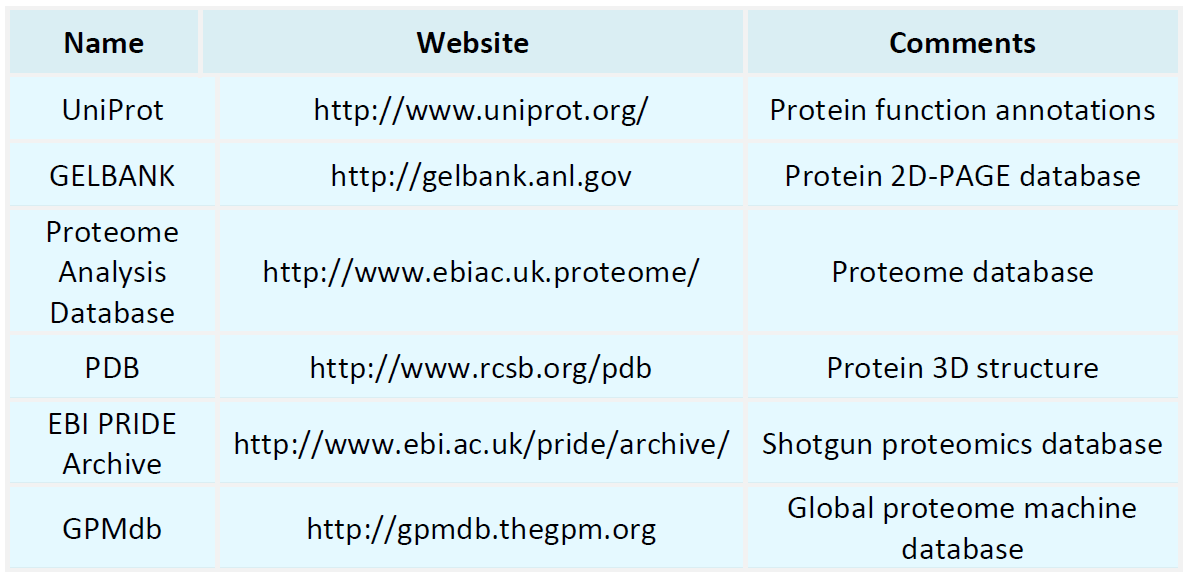
Metabolomics Databases
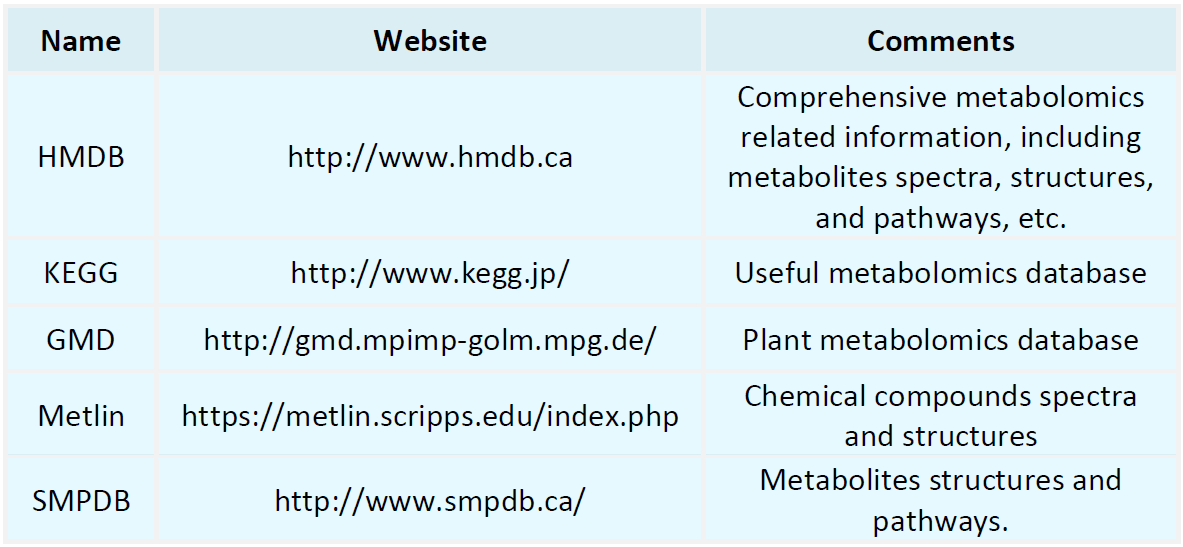
-
• Beginner’s Guide to Acetylproteomics Research: Sample Prep and Analysis Tips
As a major type of protein post-translational modification (PTM), acetylation is extensively involved in fundamental biological processes, including cellular metabolism, signal transduction, and gene expression regulation. In particular, in studies of histone regulation, mitochondrial function, and metabolic reprogramming, protein acetylation has emerged as a cutting-edge area for elucidating disease mechanisms and identifying potential therapeutic targets. With advances in high-resolution mass spectr......
-
• Mechanisms and Pathways of Protein Ubiquitination
Within complex cellular systems, the dynamic regulation of protein function represents a fundamental mechanism for maintaining cellular homeostasis. Protein ubiquitination, a highly conserved and reversible post-translational modification, is broadly involved in diverse biological processes, including cell cycle regulation, stress responses, DNA repair, and signal transduction. By covalently tagging substrate proteins, ubiquitination governs their stability, activity, subcellular localization, and int......
-
• How to Enrich Golgi Proteins for Proteomic Analysis?
The Golgi apparatus, a central membranous organelle within cells, plays essential roles in protein modification, sorting, and transport. It is critical for cellular signaling, the secretory system, and understanding disease mechanisms. Therefore, precise identification and quantitative analysis of Golgi proteins are crucial for elucidating cellular functions and disease processes. However, due to the low abundance of Golgi proteins within total cellular proteins, their enrichment and isolation remain ......
-
• How to Enrich Lactylated Peptides for Mass Spectrometry?
Lactylation is an emerging post-translational modification, first identified in 2019, which modulates protein structure and function by introducing a lactyl group (-CO-CH₃-OH) onto lysine residues. Lactylation plays critical roles in regulating carbohydrate metabolism, inflammatory responses, and tumorigenesis. For instance, histone lactylation can modulate chromatin accessibility, thereby influencing gene expression, while non-histone lactylation participates in the regulation of cellular metabolic p......
-
• Can Histone Kbu Serve as a Novel Epigenetic Biomarker?
In life science research, epigenetics has emerged as a central field for elucidating mechanisms underlying gene expression regulation. In recent years, it has been demonstrated that diverse post-translational modifications of histones not only modulate chromatin architecture but also play essential roles in gene activity, cell fate determination, and disease progression. Among these modifications, histone lysine butyrylation (Kbu, Lysine Butyrylation), as a newly identified acyl modification, has attr......
-
• How to Detect Histone Malonylation Modifications?
Histones, as fundamental components of chromatin, play a central role in the regulation of gene expression and cell fate determination through a variety of post-translational modifications. In addition to classical modifications such as acetylation, methylation, and phosphorylation, histone malonylation has emerged in recent years as an important focus in epigenetic research. This modification involves the covalent attachment of a malonyl group to lysine residues, thereby influencing chromatin accessi......
-
• Untargeted Metabolomics Analysis: Principles, Applications, and Workflow
In the context of the continuous evolution of systems biology, metabolomics has emerged as a key discipline for interpreting dynamic changes in biological processes. Among its subfields, untargeted metabolomics plays a central role in both basic research and translational medicine due to its capability to perform global, unbiased profiling of metabolites without predefined targets. Compared with targeted approaches, untargeted strategies emphasize discovery and enable the capture of potentially critic......
-
• How to Map the Golgi Proteome Using Mass Spectrometry?
In cell biology, the Golgi apparatus is a central organelle involved in protein processing, modification, and sorting, and its dysfunction is closely associated with a range of diseases, including neurodegenerative disorders and cancer. In recent years, rapid advances in mass spectrometry and proteomics have enabled systematic characterization of Golgi protein composition and its dynamic alterations at a systems level. Significance of Golgi Proteome Research The Golgi apparatus not only participates ......
-
• How to Perform Golgi Quantitative Proteomics Analysis?
The Golgi apparatus is a key membranous organelle responsible for protein modification, sorting, and trafficking, serving as an indispensable central hub for maintaining normal cellular functions. With the rapid advancement of proteomics technologies, researchers can not only identify the protein composition of the Golgi apparatus but also quantitatively characterize its dynamic alterations under different physiological or pathological conditions. Golgi quantitative proteomics provides a systematic fr......
-
• How to Isolate and Enrich Exosomes for Proteomics Analysis?
In life science research, exosomes serve as important mediators of intercellular communication and have attracted considerable attention as sources of disease biomarkers, drug delivery vehicles, and tools for basic biological research due to their cargo of proteins, nucleic acids, and lipids. For proteomics analysis, the isolation of exosomes with high purity and effective enrichment is a critical prerequisite. Overview of Exosomes and Their Proteomic Value Exosomes are nanoscale vesicles with diamet......
How to order?







