Resources
Proteomics Databases
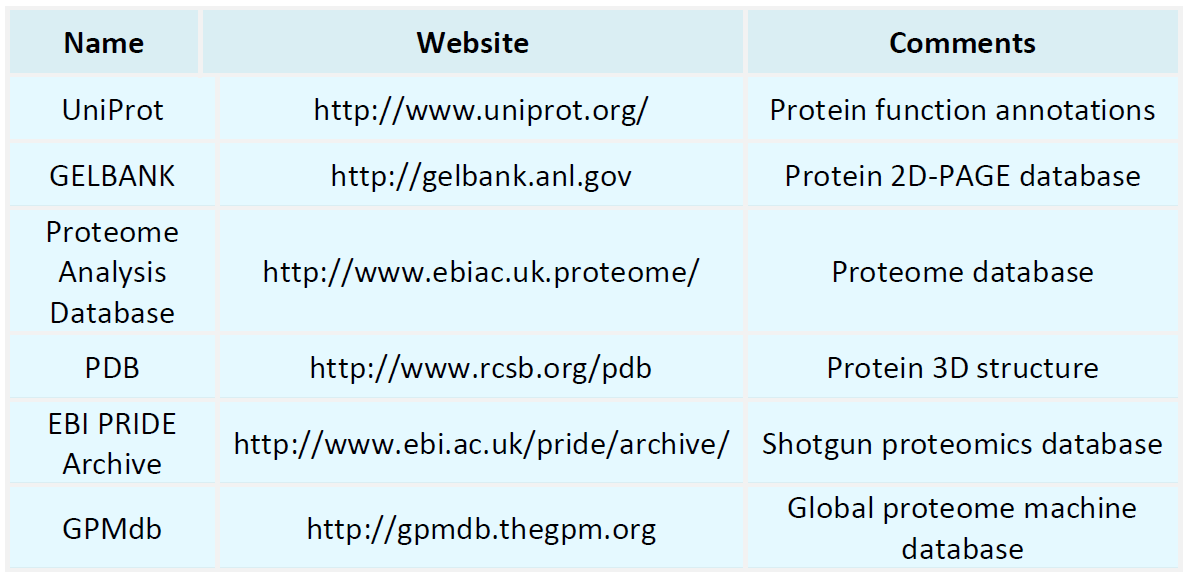
Metabolomics Databases
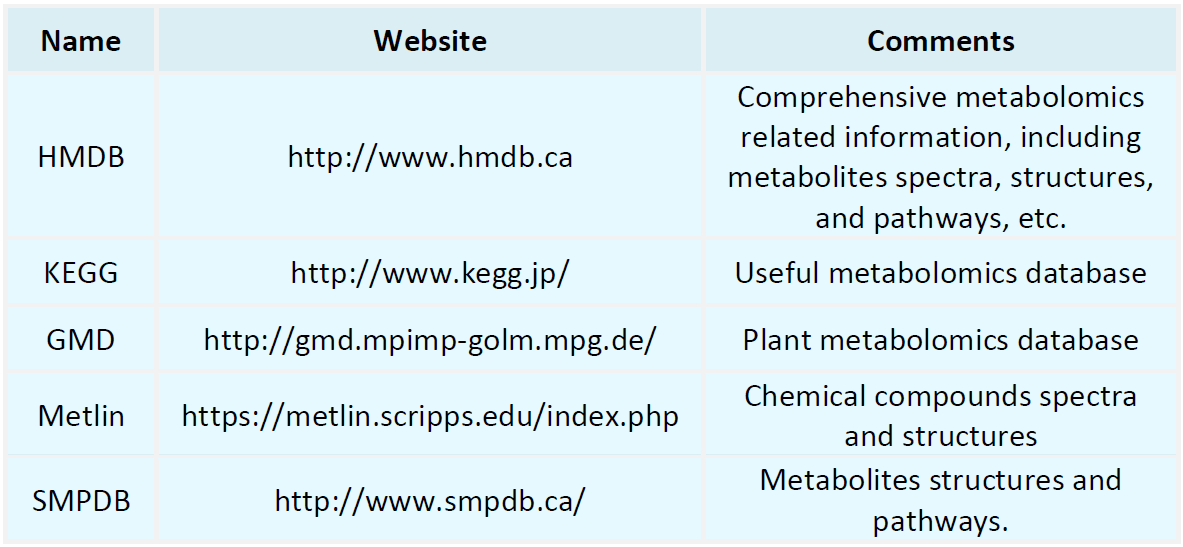
-
• How to Isolate Golgi Fractions for Subcellular Proteomics?
In subcellular proteomics, the Golgi apparatus has long attracted significant attention as a central hub for intracellular protein processing and sorting. Its essential roles in protein glycosylation, vesicular trafficking, and the secretory pathway make it a critical target for elucidating disease mechanisms and identifying potential therapeutic targets. However, precise isolation of the Golgi apparatus remains technically challenging because of its highly dynamic architecture and its extensive physi......
-
• Histone Methylation Quantitative Proteomics by LC-MS/MS
Quantitative LC-MS/MS analysis of histone methylation sites including mono-, di-, and tri-methyl states with antibody enrichment.
-
• Quantitative Enzyme Activity Analysis by ABPP and LC-MS/MS
Quantify active enzymes with activity-based probes, enrichment, and LC-MS/MS using TMT or label-free workflows.
-
• Label-Free Quantitative Proteomics for Protein Interactions
Label-free LC-MS/MS quantification applied to Co-IP and interaction proteomics without isotope tags.
-
• Absolute vs Relative Quantitative Proteomics Explained
Compare absolute and relative quantitative proteomics: isotope standards, SWATH, label-free, and TMT/iTRAQ methods.
-
• How to Use GFP Tags for Co-Immunoprecipitation?
GFP-tag Co-IP using GFP-Trap beads, gentle lysis, wash optimization, and optional LC-MS/MS for interactome discovery.
-
• Chemoproteomics: Principles, Workflow, and Applications
Chemoproteomics uses activity-based probes and LC-MS to map protein–small-molecule interactions, targets, and enzyme activity.
-
• Histone Phosphorylation in Epigenetic Regulation
Definition and roles of histone phosphorylation in transcription, DNA damage response, and mitosis, plus MS detection approaches.
-
• How to Analyze Histone Phosphorylation Using Mass Spectrometry
LC-MS/MS workflow for histone phosphorylation: acid extraction, propionylation, phosphopeptide enrichment, and site localization.
-
• Acetylation Site Analysis by Acetyl-Proteomics LC-MS/MS
Workflow for acetylation site analysis: enrichment, LC-MS/MS, and site localization by acetyl-proteomics.
How to order?







