Resources
Proteomics Databases
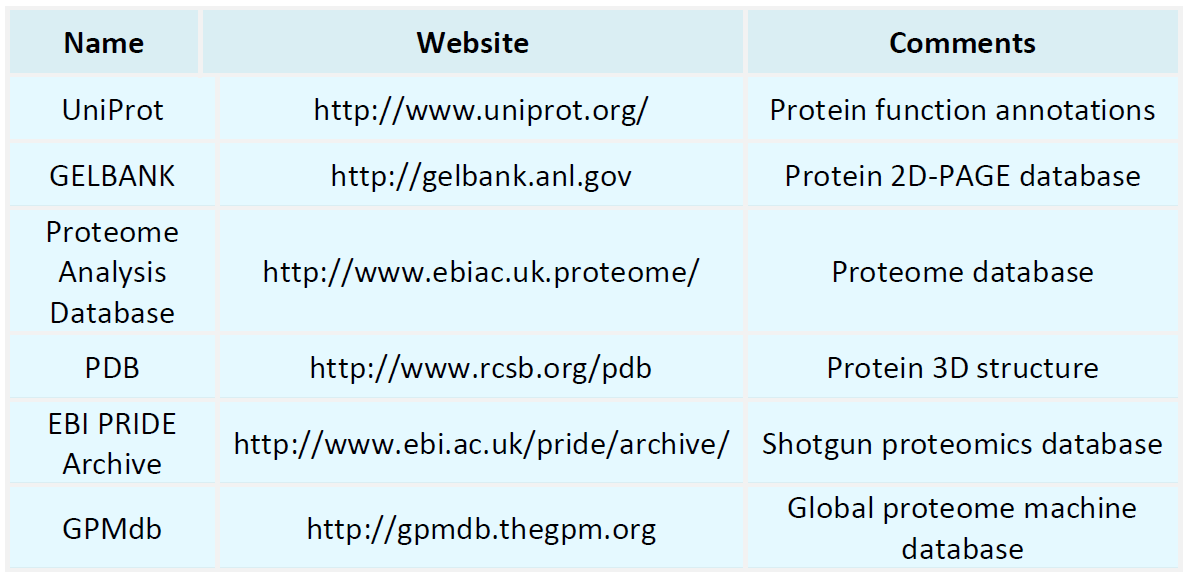
Metabolomics Databases
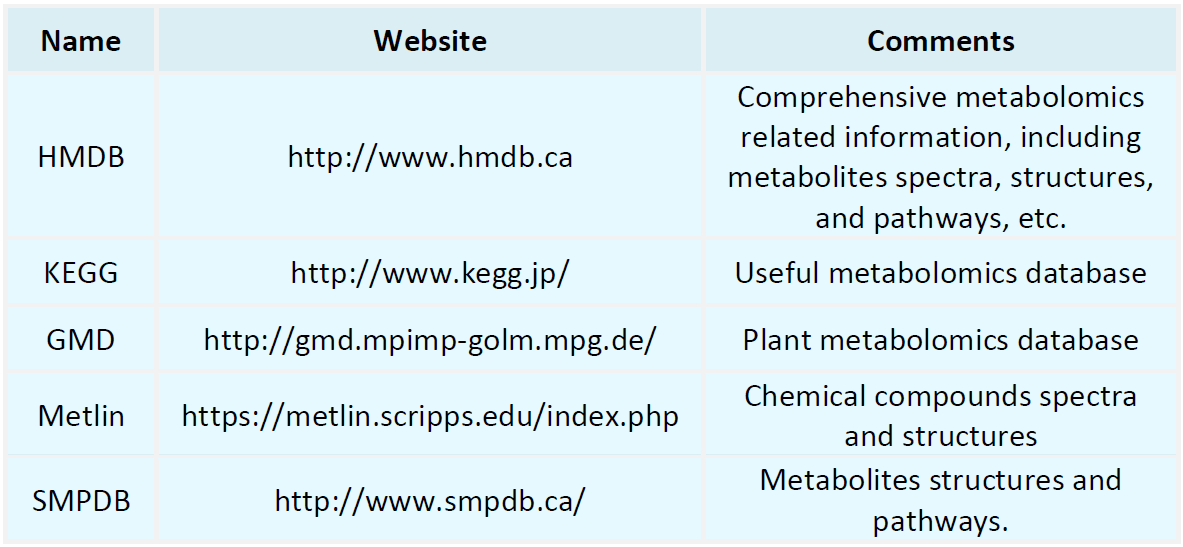
-
• What Organelle Proteomics Techniques Are Used in Modern Cell Biology?
In modern cell biology research, organelle proteomics has emerged as a powerful approach for investigating subcellular functional organization, protein localization, and intracellular signaling pathways. By leveraging high-throughput mass spectrometry technologies, researchers can systematically identify and quantify organelle-specific proteomes, thereby uncovering their dynamic regulation and functional specialization. This article provides a comprehensive overview of the major organelle proteomics t......
-
• Advantages of ESI-MS in Peptide Drug Molecular Weight Analysis
With the continuous advancement of biopharmaceutical technologies, peptide drugs, characterized by high specificity, favorable biocompatibility, and relatively low immunogenicity, have emerged as an important therapeutic strategy for cancer, autoimmune disorders, and metabolic diseases. In the development of peptide-based therapeutics, molecular weight analysis represents a fundamental and indispensable step. Owing to their structural complexity, peptide molecules are susceptible to interference from ......
-
Molecular weight represents one of the most fundamental and critical physicochemical parameters of biomolecules. Across diverse life science disciplines, including protein engineering, antibody drug development, synthetic compound validation, and metabolomics, accurate molecular weight determination serves not only as a prerequisite for structural characterization but also as a cornerstone for quality control and functional investigation. Liquid chromatography–mass spectrometry (LC-MS), characterized ......
-
High-resolution mass spectrometry (HRMS), owing to its exceptional mass accuracy, high sensitivity, and broad measurable mass range, has become an essential analytical tool in biopharmaceutical research and development. Beyond precise molecular weight determination of complex proteins, HRMS enables the resolution of subtle mass differences, providing critical support for structural characterization, impurity profiling, and quality control of biopharmaceutical products. Below, different categories of b......
-
• Acetylated Peptide Enrichment Analysis
Lysine acetylation is a prevalent post-translational modification (PTM) observed in both eukaryotic and prokaryotic organisms. It participates in epigenetic regulation, cell cycle progression, metabolic pathways, and signal transduction. In particular, histone acetylation plays a critical role in chromatin remodeling and transcriptional regulation, where the acetylation states of histones directly influence gene expression. However, compared with PTMs such as phosphorylation and ubiquitination, system......
-
• Label-Free Quantitative Glycoproteomics Using DIA-MS
Glycoproteins participate extensively in key biological processes such as cell recognition, signal transduction, and immune regulation, and dynamic alterations in their glycosylation states are often closely associated with various diseases. Glycoproteomics has therefore emerged as a crucial approach for biomarker discovery and mechanistic studies. To enable systematic quantification of glycosylated proteins, more stringent analytical requirements have been placed on glycoprotein analysis workflows. D......
-
• Comprehensive Workflow for Phosphoproteomics Data Processing and Bioinformatic Interpretation
Phosphoproteomics, a high-throughput mass spectrometry-based analytical approach for characterizing protein phosphorylation, provides a crucial means to interrogate the regulatory roles of phosphorylation in cellular signaling, cell-cycle control, and metabolic regulation. As a dynamic and reversible post-translational modification (PTM), phosphorylation modulates a broad spectrum of biological processes and has become indispensable in mechanistic studies of signal transduction and disease-relevant pa......
-
• Principles and Biological Significance of Protein Lactylation Modification
Post-translational modification (PTM) is a core regulatory mechanism governing protein function, subcellular localization, and molecular interactions. In addition to classical modifications such as phosphorylation, acetylation, and ubiquitination, an emerging modification - lysine lactylation (Kla) - has been recently uncovered. First reported by Zhang et al. in 2019, lactylation is derived from lactate generated during cellular metabolism, a metabolite traditionally viewed as a terminal product of gl......
-
• Protein Quantification Technology‑TMT Labeling Quantitation
In modern life science research, accurate protein quantification plays a central role in elucidating biological mechanisms, identifying disease biomarkers, and advancing drug development. Compared with label-free quantification approaches, Tandem Mass Tag (TMT)-based quantification has emerged as one of the most widely adopted strategies in proteomics due to its high throughput, multiplexed sample analysis, and reduced inter-batch variability. This article provides a comprehensive overview of TMT labe......
-
• Co‑immunoprecipitation (Co‑IP) Overview
Protein-protein interactions form the molecular basis of cellular signal transduction, metabolic regulation, and disease pathogenesis. As a classical approach for investigating endogenous protein interactions, co-immunoprecipitation (Co-IP) remains widely applied in molecular biology and proteomics research owing to its operational robustness, high specificity, and broad applicability. In particular, Co-IP is well suited for validating interactions between defined protein pairs. What Is Co-immunoprec......
How to order?







