Resources
Proteomics Databases
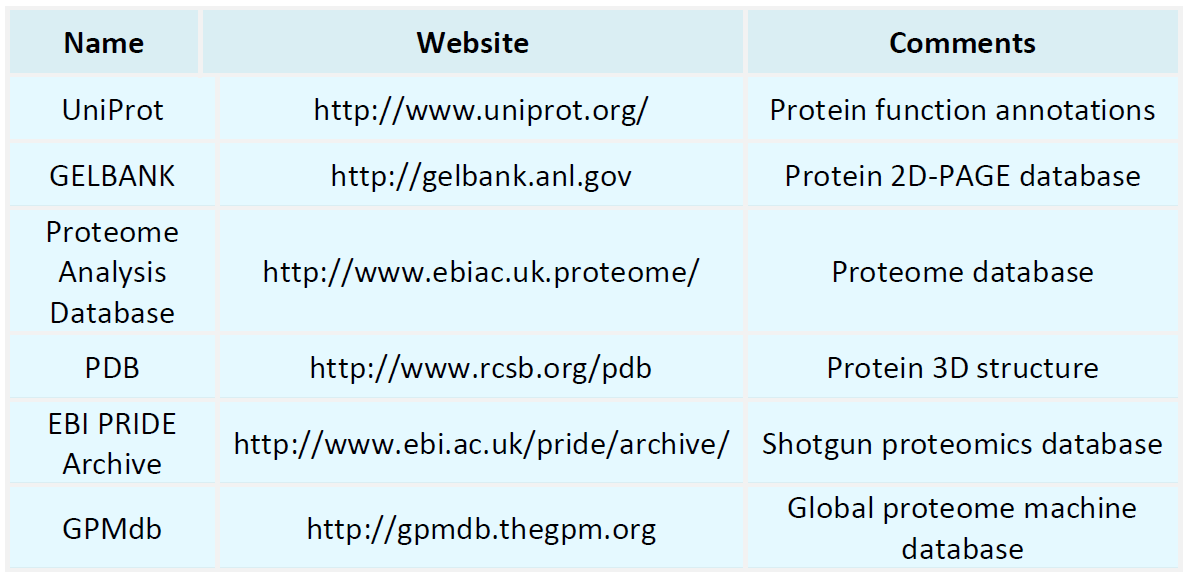
Metabolomics Databases
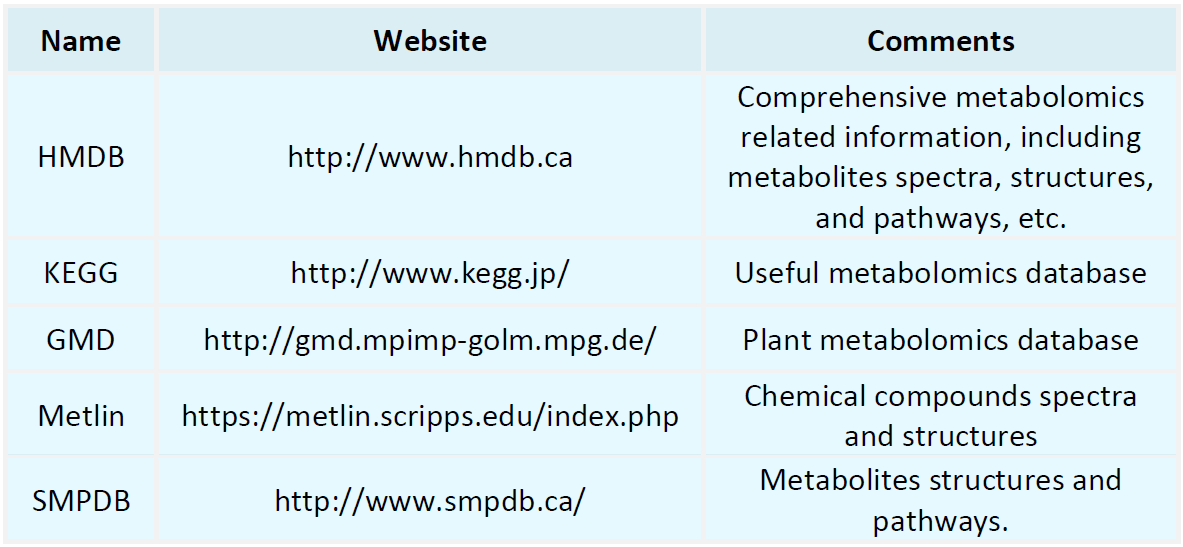
-
• How to Improve Reliability and Reproducibility of PPI Experimental Results?
Protein-protein interactions (PPIs) play fundamental roles in cellular signaling, metabolic regulation, and disease pathogenesis. With the rapid advancement of omics technologies, PPI research has become a central focus in life science studies. However, due to the intrinsic complexity of experimental systems and pervasive non-specific interactions, PPI experiments frequently suffer from poor reproducibility and elevated false-positive rates. Enhancing experimental reliability and reproducibility has t......
-
• How to Normalize Data in Label-Free Quantitative Proteomics?
In label-free quantitative proteomics (LFQ), data normalization represents a fundamental step for ensuring inter-sample comparability and minimizing technical variability. In the absence of appropriate normalization, technical noise, such as differences in sample loading amounts and instrument performance drift, can obscure genuine biological variation, thereby introducing bias into the identification of differentially expressed proteins. Why Is Normalization Necessary for Label-Free Quantitative Pro......
-
• How to Interpret Peptide Coverage and Protein Scores in Shotgun Data?
Shotgun proteomics typically yields hundreds to thousands of identified proteins and peptide-spectrum matches (PSMs). However, determining which identifications are reliable, which proteins warrant downstream investigation, and which datasets are suitable for differential analysis or functional annotation remains a critical analytical challenge. Among the metrics used to assess identification confidence, peptide sequence coverage and protein-level scoring represent two key parameters that should be ev......
-
• Critical Roles of Histone PTMs in Cancer Epigenetics
Tumor initiation and progression are not solely driven by genetic mutations; instead, deeper layers of regulation frequently reside in epigenetic mechanisms. As fundamental structural components of chromatin, histones undergo diverse post-translational modifications (PTMs), including methylation, acetylation, and phosphorylation. Together, these modifications constitute a complex epigenetic code that precisely governs chromatin accessibility and transcriptional states. In cancer cells, histone PTMs ar......
-
• Dynamic Changes of Histone PTMs During Embryonic Development
Embryonic development is a highly coordinated biological process characterized by the precise spatiotemporal regulation of thousands of genes. Although the genomic DNA sequence within the nucleus remains largely unchanged throughout development, cellular identities progressively diverge along distinct differentiation trajectories. A critical regulatory layer governing whether genes remain transcriptionally silent or become activated is provided by chromatin-based epigenetic mechanisms, particularly hi......
-
• From Sample Preparation to Data Interpretation: A Complete Workflow for Histone PTMs Analysis
Histone post-translational modifications (PTMs) represent critical regulatory mechanisms governing chromatin organization and gene expression. These modifications play essential roles in a wide range of biological processes, including embryonic development, cell differentiation, and DNA damage repair. With the rapid advancement of high-resolution mass spectrometry technologies, it has become possible to systematically profile histone modification landscapes at a global scale, providing robust technica......
-
• Structural Role and Stability of Disulfide Bonds in Proteins
Within the intricate landscape of protein three-dimensional architecture, disulfide bonds represent a prominent structural constraint. Formed as covalent linkages between cysteine residues, disulfide bonds play essential roles in maintaining higher-order protein structures, enhancing conformational stability, and modulating functional activity. In particular, in extracellular proteins, membrane proteins, and antibody-based therapeutics, disulfide bonds are often indispensable for proper folding and bi......
-
• The Critical Role of Protein-Protein Interactions in Drug Target Discovery
In conventional drug discovery pipelines, target identification has traditionally focused on individual proteins or discrete signaling pathways. However, advances in systems biology and high-throughput omics technologies have fundamentally reshaped this perspective. It is now widely recognized that most diseases do not arise from the dysfunction of isolated proteins, but rather from perturbations within complex intracellular protein-protein interaction (PPI) networks. Consequently, systematic characte......
-
• Troubleshooting Common Issues in iTRAQ Proteomics Experiments
iTRAQ (Isobaric Tags for Relative and Absolute Quantitation) is widely employed in quantitative proteomics and enables the relative quantification of up to 8 or 16 samples within a single mass spectrometry run. This capability makes it particularly suitable for comparative analyses involving multiple samples, such as clinical cohorts, disease models, and studies of drug mechanisms of action. Nevertheless, the high throughput and sensitivity of iTRAQ-based workflows place stringent demands on experimen......
-
Proteins serve as the primary functional units within cells. However, individual proteins rarely operate in isolation; instead, most biological processes are executed through coordinated protein-protein interactions (PPIs). These interactions are fundamental to diverse biological processes, including signal transduction, metabolic regulation, and immune responses, and they also play pivotal roles in the initiation and progression of diseases. Comprehensive experimental identification of all possible P......
How to order?







