Resources
Proteomics Databases
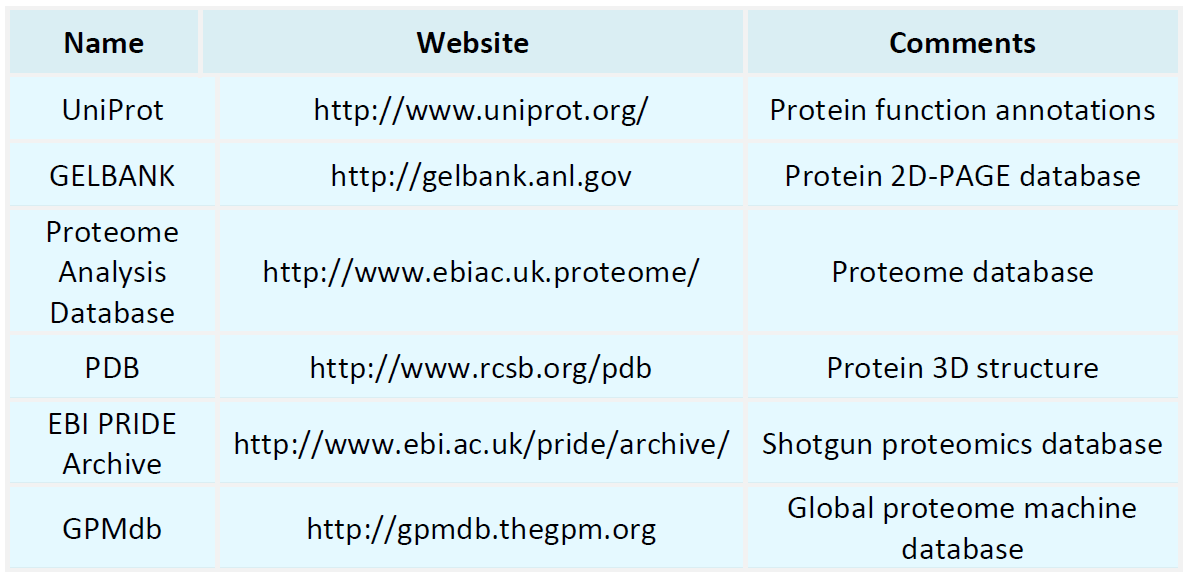
Metabolomics Databases
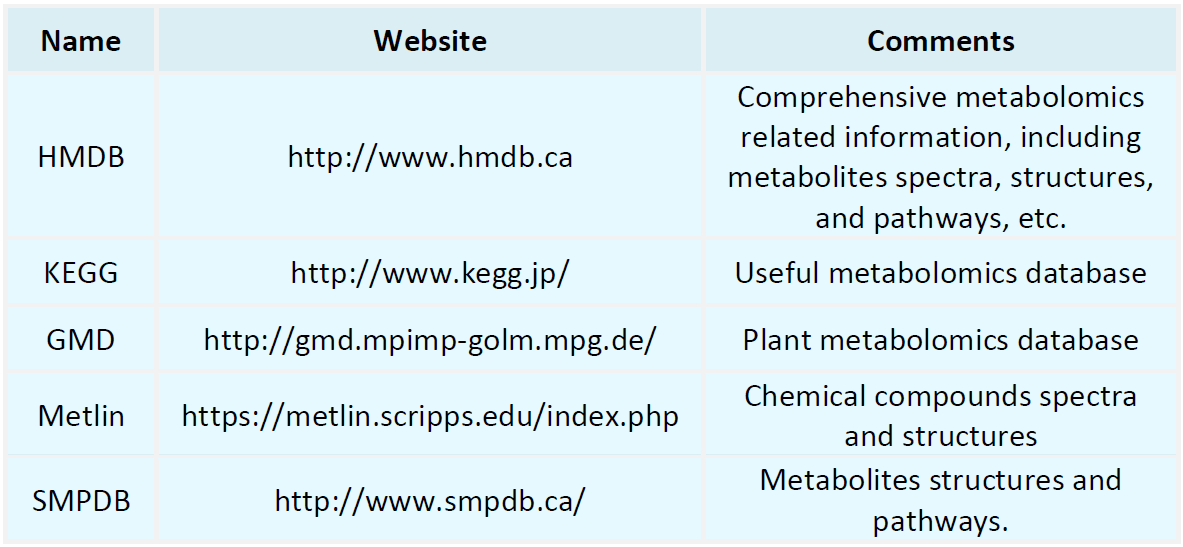
-
• How to Perform Qualitative Shotgun Protein Identification Using DDA Mode?
In life science research, proteins serve as a central link between genomic information and cellular functions. To systematically identify and characterize the full complement of proteins in biological samples, shotgun protein identification has become one of the most widely adopted strategies in proteomics. This approach involves the direct introduction of enzymatically digested peptide mixtures into a mass spectrometry system, coupled with data-dependent acquisition (DDA) for high-throughput data col......
-
• Comprehensive Guide to 4D Label-Free Quantitative Proteomics Workflow
In the post-genomic era, proteomics has emerged as a pivotal approach for elucidating biological processes, uncovering disease mechanisms, and discovering biomarkers. Specifically, label-free quantitative proteomics (LFQ) has gained widespread application in both basic and translational biomedical research due to its high throughput and the advantage of eliminating the need for costly labeling reagents. With the rapid advancement of mass spectrometry technologies, 4D-based label-free quantification pl......
-
• Principles of Top-Down Proteomics
Top-Down proteomics is a mass spectrometry-based analytical strategy that directly analyzes intact protein molecules. Its core principle is to subject proteins to mass spectrometric fragmentation and sequencing without prior enzymatic digestion, thereby enabling accurate protein identification, detection of post-translational modifications (PTMs), and recognition of protein variants. Compared with Bottom-Up proteomics, the Top-Down approach allows more comprehensive protein identification and structur......
-
• How to Integrate Shotgun Proteomics and Targeted Proteomics for Joint Analysis?
In life science research, proteins are central executors of biological function. Their expression levels, modification states, and interaction networks often directly shape cellular fate and disease progression. To comprehensively characterize these dynamic changes, a range of mass spectrometry–based approaches has been developed. Among the most widely used are Shotgun proteomics (data-dependent acquisition, DDA) and targeted proteomics (e.g., PRM/MRM). Nevertheless, each approach has notable limitati......
-
• Histone Modifications in Health and Disease
The Human Genome Project delineated the human genetic blueprint; however, DNA sequence information alone is insufficient to account for the complexity of cell fate determination. Cells that share an identical genome can nonetheless perform markedly distinct functions across tissues, a divergence that is largely attributable to epigenetic regulation. Within this regulatory framework, histone modifications constitute a central and highly dynamic mechanism that shapes gene expression, chromatin organizat......
-
• Application of DIA Technology in Ubiquitin Proteomics
Ubiquitination is a major post-translational modification (PTM) in which the 76-amino-acid ubiquitin protein is covalently attached to lysine residues of substrate proteins. This modification regulates a wide range of cellular processes, including protein stability, signal transduction, and DNA repair. Conventional mass spectrometry strategies such as data-dependent acquisition (DDA) have enabled the identification of ubiquitinated peptides, yet they often suffer from sampling bias and missing values.......
-
• Advantages and Challenges of TMT Labeling for Lysine Acetylation Quantification
Lysine acetylation is a key post-translational modification (PTM) that broadly contributes to fundamental biological processes, including chromatin remodeling, metabolic regulation, and signal transduction. Although its biological roles have been progressively elucidated, robust quantitative profiling remains technically challenging because acetylation is typically low in stoichiometry, highly dynamic, and tightly coupled to tissue context. Achieving sensitive, high-throughput, and reproducible quanti......
-
• What Is Label-Free Analysis?
In multi-omics research areas such as proteomics and metabolomics, quantitative analysis is fundamental to the interpretation of biological variation. With the rapid advancement of mass spectrometry technologies, label-free analysis, commonly referred to as Label-Free Quantification (LFQ), has increasingly been adopted as a primary strategy for quantitative proteomic studies owing to its high throughput, cost efficiency, and streamlined workflow. Label-free analysis enables the relative quantification......
-
• What Is Protein-Protein Interaction (PPI) Network Visualization? Recommended Tools and Software
Protein-protein interaction (PPI) networks represent a fundamental analytical framework for elucidating cellular functions, signaling pathway regulation, and disease mechanisms. Protein-protein interaction visualization involves the graphical representation of complex interaction relationships, enabling researchers to intuitively identify key proteins, functional modules, and network topological features, thereby facilitating biological interpretation and discovery. What Is Protein-Protein Interactio......
-
• What Is High-Throughput PPI Screening and What Are the Cutting-Edge Technologies?
High-throughput protein-protein interaction (PPI) screening refers to a set of methodologies that leverage automation and large-scale experimental platforms to systematically detect and analyze interactions among large numbers of proteins within a limited time frame. This strategy is essential for protein function annotation, signaling pathway elucidation, investigation of disease mechanisms, and drug target discovery, and represents a key enabling tool for deciphering cellular networks and biological......
How to order?







