Resources
Proteomics Databases
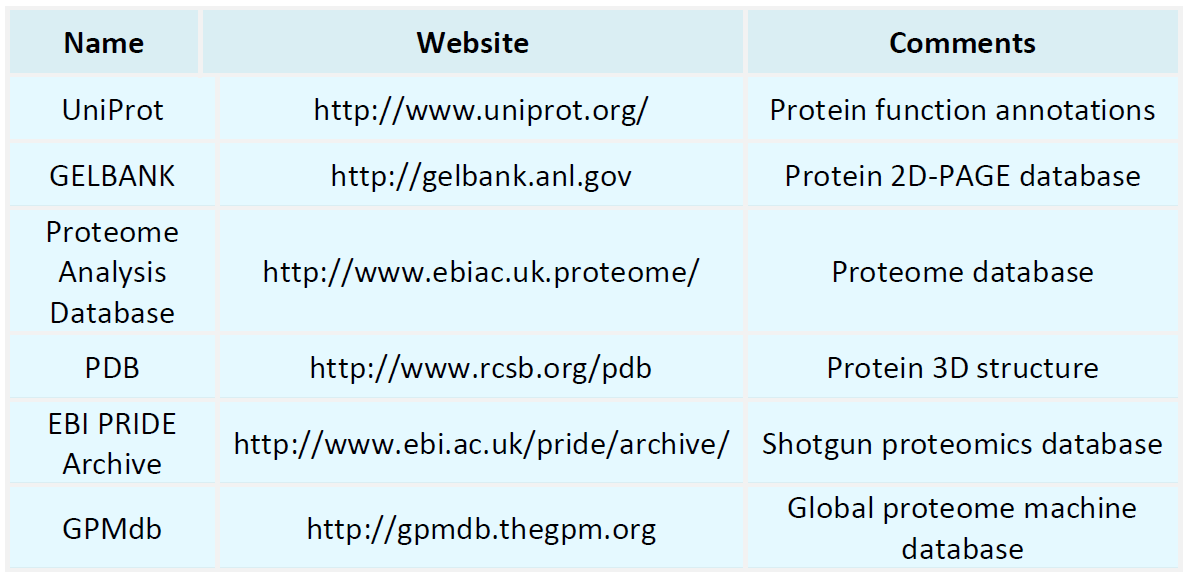
Metabolomics Databases
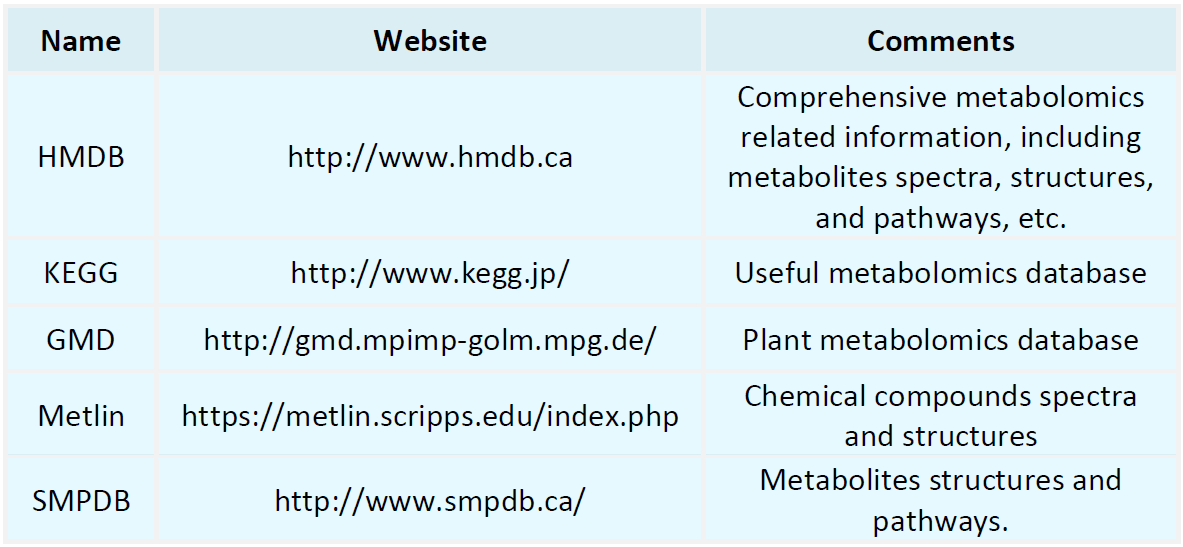
-
• What Are Protein-Protein Interactions?
Protein-protein interactions (PPIs) refer to the processes by which two or more protein molecules associate to form functional complexes or regulate one another through non-covalent forces, including hydrogen bonding, hydrophobic interactions, and electrostatic forces. PPIs are ubiquitous within cells and underpin a wide range of essential biological processes. From signal transduction and cell cycle control to metabolic pathways and immune responses, virtually all cellular activities rely on precisel......
-
• How to Analyze Protein–Protein Interactions?
Protein-protein interactions (PPIs) represent fundamental mechanisms underlying cellular function and biological processes. They play essential roles in signal transduction, metabolic regulation, and gene expression. Investigating PPIs facilitates the elucidation of disease mechanisms, the identification of therapeutic targets, and the construction of regulatory networks. With advances in mass spectrometry technologies, PPI detection has progressed from conventional validation approaches to high-throu......
-
• What Is Label-Free Quantification in Proteomics?
Quantitative analysis is a fundamental approach in proteomics research for elucidating dynamic biological processes, identifying differentially expressed proteins, and investigating disease mechanisms. Compared with labeling-based quantification strategies, such as SILAC, iTRAQ, and TMT, label-free quantification (LFQ) has been increasingly adopted in recent years owing to its operational simplicity, broad applicability, and relatively low cost. In particular, LFQ offers distinct advantages for the an......
-
• The Role of Phosphoproteomics in Multi-Omics Integration
In the post-genomic era, life science research is transitioning from isolated, single-layer investigations toward comprehensive systems-level integration. Technologies such as transcriptomics, proteomics, metabolomics, epigenomics, and single-cell omics each provide snapshots of biological systems at distinct molecular layers. However, only through multi-omics integration can the dynamic and coordinated nature of biological processes be fully reconstructed. Within this integrative framework, phosphopr......
-
• Common Pitfalls in Phosphoproteomics Experiments And How to Avoid Them
Phosphoproteomics is a critical research field dedicated to the systematic investigation of protein phosphorylation and its functional roles in cellular biology. Phosphorylation, dynamically regulated by protein kinases and phosphatases, modulates protein activity, subcellular localization, stability, and molecular interactions. This post-translational modification plays a central role in a wide range of biological processes, including signal transduction, cell cycle control, metabolic regulation, gen......
-
• Phosphorylation Analysis by Mass Spectrometry
Protein phosphorylation is one of the most prevalent and functionally important post-translational modifications in cells, with approximately one-third of eukaryotic proteins undergoing phosphorylation. By modulating protein activity, stability, conformation, and intermolecular interactions, phosphorylation serves as a central mechanism in cellular signal transduction, metabolic regulation, cell cycle control, and stress responses. Accurate identification and quantification of phosphorylation events a......
-
• Comprehensive Overview of Mass Spectrometry-Based Protein Phosphorylation Analysis Technologies
Protein phosphorylation is one of the most prevalent and functionally significant post-translational modifications, playing essential roles in a wide range of biological processes, including cell proliferation, differentiation, metabolism, and apoptosis. Precise identification and quantitative characterization of phosphorylation sites enable researchers to systematically decipher the dynamic regulation of cellular signaling networks, elucidate molecular mechanisms underlying disease pathogenesis, and ......
-
• How to Integrate Spatial Proteomics with Multi-Omics Approaches?
The rapid advances in single-cell omics, spatial transcriptomics, and high-throughput mass spectrometry have propelled life-science research into an era of high-dimensional data integration. Conventional proteomics captures changes in protein abundance, yet it often cannot address a fundamental question: where, precisely, are these proteins located within tissues or cells? Spatial proteomics has been developed to fill this gap by enabling visualization and quantitative profiling of protein distributio......
-
• Common Types of Histone Modifications and Their Mass Spectrometric Signatures
Histones are core structural components of chromatin that not only facilitate DNA packaging and maintain chromatin architecture, but also play essential regulatory roles in gene expression, DNA repair, and chromatin remodeling through a wide range of post-translational modifications (PTMs). A broad spectrum of histone modifications has been identified, among which acetylation, methylation, phosphorylation, and ubiquitination are the most extensively studied. Collectively, these modifications constitut......
-
• LC-MS/MS-Based Quantitative Ubiquitinomics
Ubiquitination is a critical post-translational protein modification involved in cellular regulation and participates in numerous essential physiological processes, including protein degradation, signal transduction, cell cycle control, and DNA damage response. However, the accurate and sensitive quantification of ubiquitinated proteins in complex biological systems remains highly challenging. LC-MS/MS (liquid chromatography–tandem mass spectrometry) has become a core analytical platform for the quant......
How to order?







