Resources
Proteomics Databases
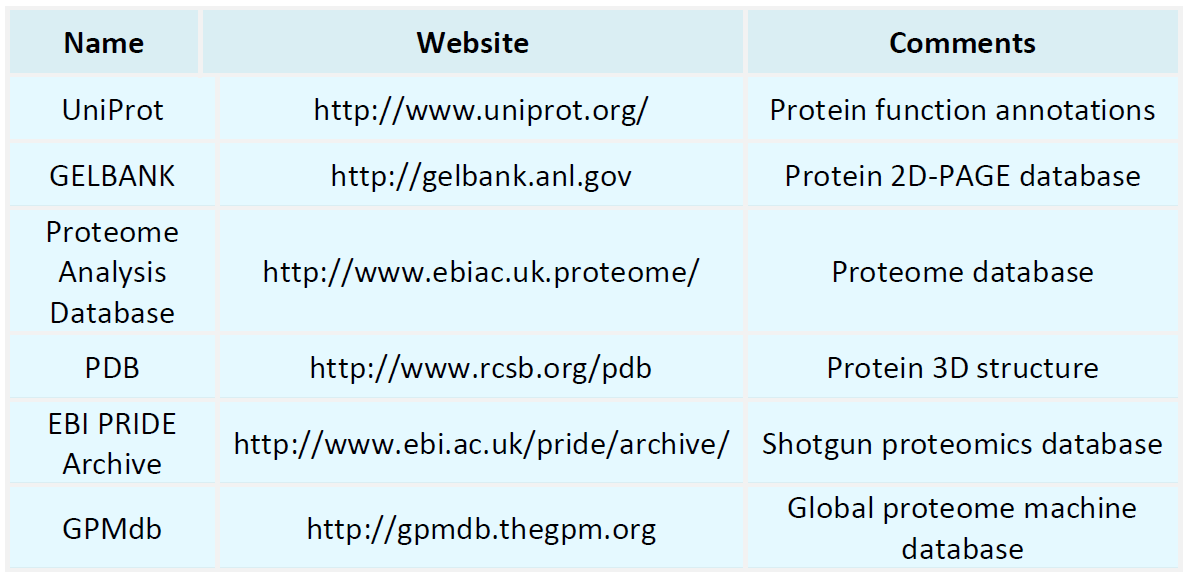
Metabolomics Databases
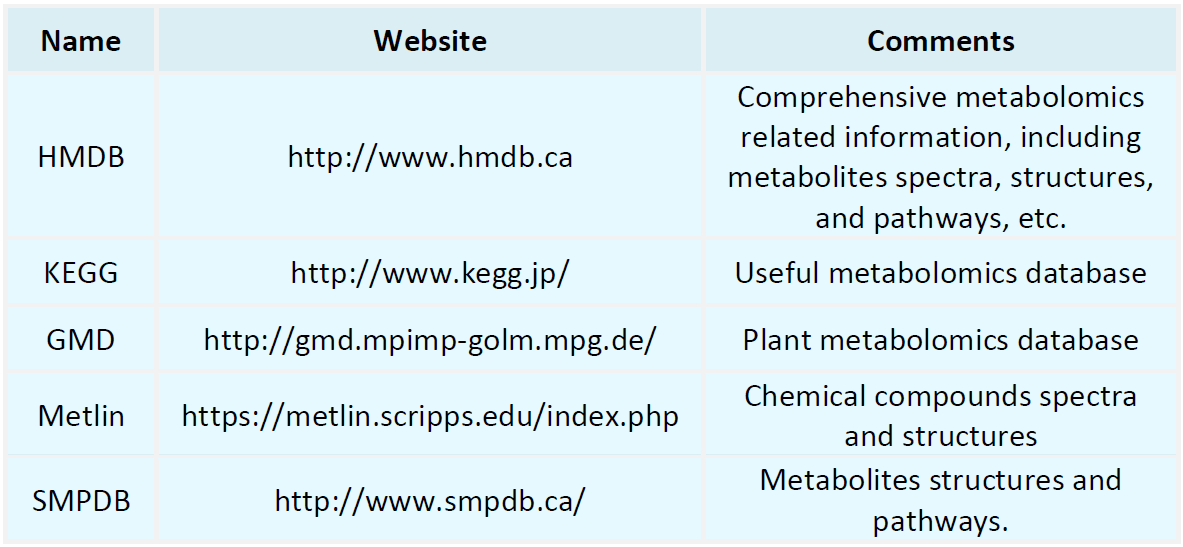
-
Compare six leading bottom-up proteomics data analysis tools for identification, quantification, PTM analysis, and targeted workflows.
-
Explore how bottom-up proteomics supports clinical research through biomarker discovery, disease subtyping, drug mechanism studies, and QC-aware workflows for plasma, tissue, and biofluid samples.
-
• Principles of Quantitative Ubiquitinomics: Enrichment, LC-MS/MS, and Quantification Strategies
Learn the core principles of quantitative ubiquitinomics, including diGly peptide enrichment, TMT and SILAC quantification, LC-MS/MS site mapping, and analysis of ubiquitination dynamics.
-
A practical LC-MS/MS workflow for histone β-hydroxybutyrylation (Kbhb) analysis covering histone extraction, digestion, peptide enrichment, acquisition settings, site localization, and quantification.
-
• Step-by-Step ABPP Experiment Protocol: Probe Design, Labeling, LC-MS/MS, and Activity Profiling
A practical step-by-step ABPP protocol covering research objectives, activity-based probe selection, in vitro and in situ labeling, enrichment, LC-MS/MS acquisition, and activity profile analysis.
-
A practical guide to how activity-based protein profiling (ABPP) supports high-throughput compound screening through competitive labeling, functional target validation, off-target profiling, and LC-MS/MS readouts.
-
A practical guide to protein detection methods used with SDS-PAGE, including Western blot, mass spectrometry, 2D-PAGE, in-gel workflows, method selection, and common limitations.
-
A practical guide to efficient and accurate protein sequencing methods, comparing mass spectrometry, Edman degradation, de novo interpretation, terminal analysis, and gene-based sequence inference for research and biopharma workflows.
-
A practical guide to glycosylation site analysis in protein function studies, covering N- and O-glycosylation, LC-MS/MS workflows, enrichment, site occupancy, glycoform interpretation, and disease or biopharma applications.
-
A practical guide to cell surface proteomics analysis, covering biotinylation, shaving, membrane protein enrichment, DDA and DIA workflows, localization filtering, and biomarker discovery.
How to order?







