Resources
Proteomics Databases
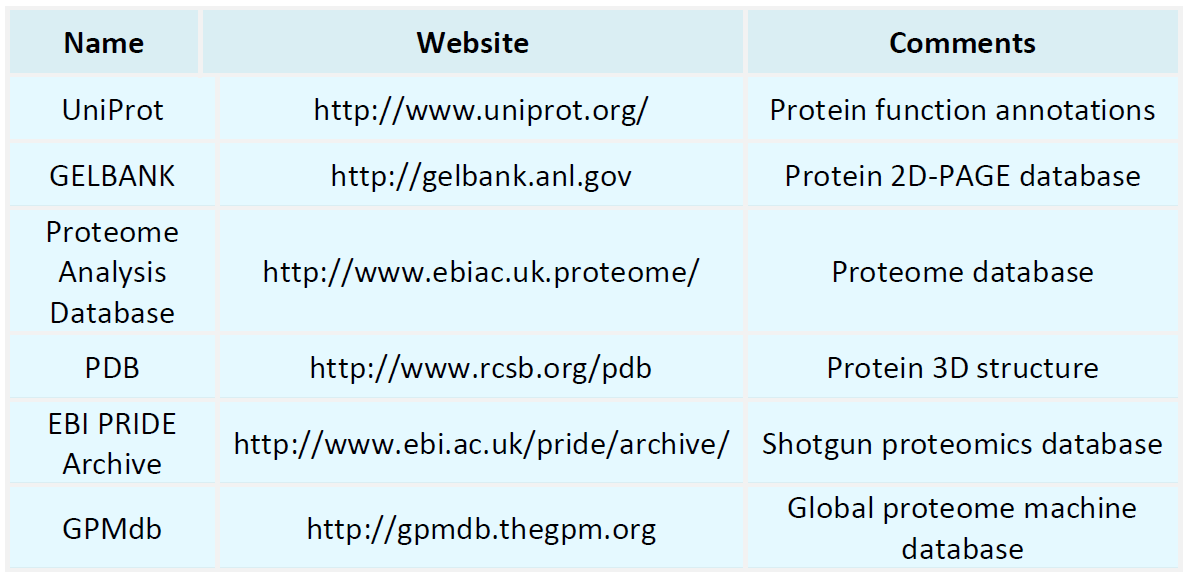
Metabolomics Databases
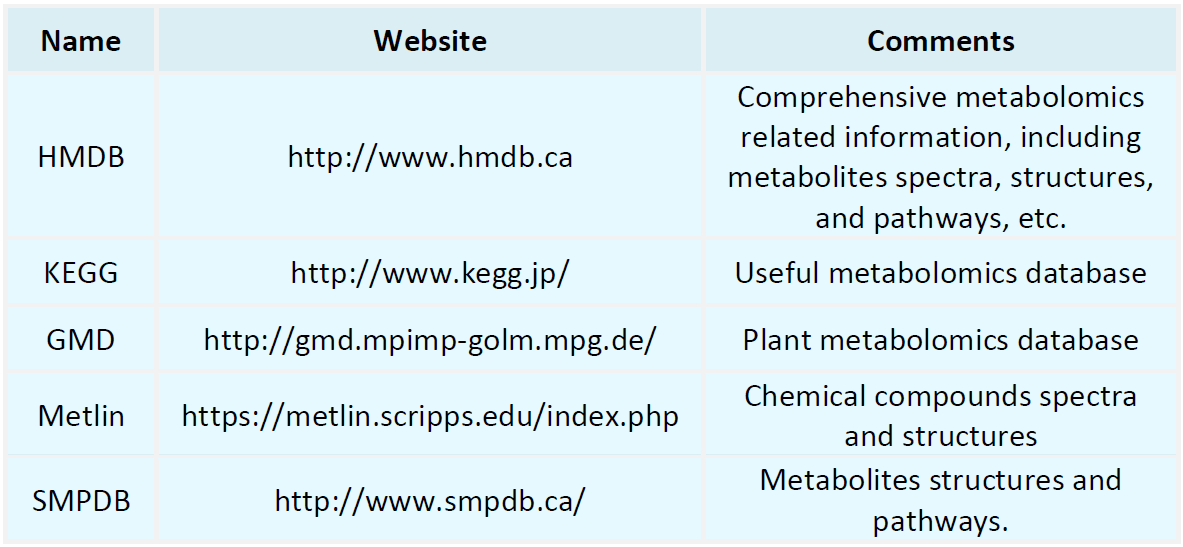
-
• Integrated Solutions for Protein Structure Identification and Functional Prediction
In the post-genomic era, elucidating the three-dimensional structure and functional properties of proteins is a critical step toward advancing our understanding of biological processes. Proteins are fundamental functional units of cells, and the characterization of their structures and functions forms the essential basis for drug discovery, the investigation of disease mechanisms, and precision medicine. Classical structural biology techniques, such as X-ray crystallography, nuclear magnetic resonance .....
-
• How Can Protein Structure Analysis Facilitate Biopharmaceutical Development?
In the rapidly evolving field of modern biopharmaceutical research and development, proteins, whether as primary therapeutic targets or as drug molecules, possess structural attributes that directly determine their biological functions and mechanisms of action. Protein structure analysis, as a central approach for elucidating protein molecular conformations, plays an indispensable role throughout the entire lifecycle of biopharmaceuticals, from discovery and design to optimization and quality control.
-
• What Are the Advantages and Limitations of Circular Dichroism (CD) Spectroscopy?
Circular dichroism (CD) spectroscopy, a spectroscopic technique for investigating the conformations of biological macromolecules, has been widely employed in structural biology and biophysics owing to its ease of operation and readily interpretable results. In particular, CD spectroscopy serves as an indispensable tool in the secondary structure analysis of chiral molecules such as proteins and nucleic acids, in monitoring conformational changes, and in probing intermolecular interactions. As with all .....
-
• How Does Circular Dichroism (CD) Spectroscopy Contribute to Drug Development?
Modern drug development increasingly depends on the precise characterization of the structure, conformation, and stability of biological macromolecules. In particular, for protein therapeutics, biopharmaceuticals, and studies of small molecule–target interactions, structural information not only determines the underlying functional mechanisms but also directly influences efficacy, safety, and controllability. Owing to its rapid measurement, high sensitivity, and capability to analyze structures in ......
-
• What Can CD Spectroscopy Really Tell Us About Proteins?
In modern protein research, Circular Dichroism (CD) spectroscopy is a widely applied analytical method characterized by its operational simplicity. By detecting the differential absorption arising from the interaction between light and chiral molecules, CD spectroscopy reveals multiple critical aspects of protein structure. Although the information it provides is relatively indirect, CD spectroscopy remains indispensable for rapid structural evaluation, monitoring of conformational changes, and studies of..
-
• Near UV Circular Dichroism (CD) Spectroscopy Analysis
In structural biology, accurate characterization of the higher-order structure of proteins is essential for understanding their functions and mechanisms. Near-UV Circular Dichroism (CD) spectroscopy, a non-destructive, efficient, and solution-compatible technique, has been widely adopted in studies of protein conformation, drug screening, and quality control of biopharmaceuticals. This review provides a systematic overview of the fundamental principles, practical advantages, and critical relevance of ......
-
• Pros and Cons of 3 Common Protein Structure Analysis Methods
Proteins, as fundamental molecules driving biological processes, have their three-dimensional structures directly determining their functions. Accurate analysis of protein structure not only facilitates elucidation of biological mechanisms but also provides critical insights for drug discovery and disease mechanism studies. Currently, the principal techniques for protein structure analysis include X-ray crystallography, nuclear magnetic resonance (NMR) spectroscopy, and cryo-electron microscopy (Cryo-EM).
-
• CD-Based Peptide Secondary Structure Analysis
In life sciences, the conformational states of peptide molecules critically influence their biological functions, particularly in essential processes such as signal transduction, immune recognition, and drug development. The peptide’s secondary structure, comprising α-helices, β-sheets, and random coils, not only defines its spatial architecture but also plays a central role in molecular interactions. Circular Dichroism (CD) spectroscopy, a rapid and sensitive spectroscopic technique, has become a ......
-
• Top 5 Methods for Protein Structure Analysis and Their Applications
The function of a protein is intrinsically linked to its three-dimensional structure. Protein structure analysis not only facilitates the interpretation of biological behavior but is also extensively utilized in diverse research areas, including drug discovery, antibody engineering, and the elucidation of metabolic pathways. Currently, researchers have access to multiple analytical techniques for analyzing protein structure, each with distinct advantages, limitations, and suitability for specific protein...
-
• Protein Primary Structure Characterization Methods
Proteins are fundamental biomolecules essential for life processes, and their structures are intrinsically linked to their functions. The primary structure of a protein refers to the linear sequence of amino acids, which serves as the foundation for the formation of higher-order structures such as secondary, tertiary, and quaternary conformations. Accurate determination of a protein’s primary structure is critical for elucidating its biological functions, investigating structural alterations, developing....
How to order?







